 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Glyma.10G053500.1.p | ||||||||
| Common Name | GLYMA_10G053500, LOC100796522 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 701aa MW: 77266.4 Da PI: 7.5109 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 72.4 | 5.4e-23 | 118 | 219 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFk 83
f k+lt+sd+++ g +++p+ +ae++ ++ + + +d +g++W++++iyr++++r++lt+GW+ Fv+ ++L +gD++vF
Glyma.10G053500.1.p 118 FAKTLTQSDANNGGGFSVPRYCAETIfprldysVDPPVQ--NILAKDVHGETWKFRHIYRGTPRRHLLTTGWSTFVNHKKLVAGDSIVFL 205
89****************************996445555..69999******************************************** PP
E-SSSEE..EEEEE-S CS
B3 84 ldgrsefelvvkvfrk 99
+ +++ l+v+++r+
Glyma.10G053500.1.p 206 --RAENGDLCVGIRRA 219
..4488999****997 PP
| |||||||
| 2 | Auxin_resp | 109.4 | 3.8e-36 | 287 | 370 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrl.sGtvvgvsdldpvrWpnSkWrsLk 83
a++ a++k++FevvY+Prast+eF+vk++ ve+al+++++ G Rfkmafetedss++++ +Gt+++ + +dp++WpnS+Wr+L+
Glyma.10G053500.1.p 287 ASNLAANKKPFEVVYYPRASTPEFCVKASLVEAALQIRWCSGIRFKMAFETEDSSRISWfMGTISSAQVADPLNWPNSPWRLLQ 370
67899*****************************************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 4.3E-36 | 111 | 221 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 5.76E-29 | 112 | 219 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 4.44E-21 | 117 | 218 | No hit | No description |
| PROSITE profile | PS50863 | 12.872 | 118 | 220 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 6.9E-21 | 118 | 220 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 4.1E-21 | 118 | 219 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 1.0E-31 | 287 | 370 | IPR010525 | Auxin response factor |
| PROSITE profile | PS51745 | 17.57 | 614 | 694 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007389 | Biological Process | pattern specification process | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0048829 | Biological Process | root cap development | ||||
| GO:0051301 | Biological Process | cell division | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 701 aa Download sequence Send to blast |
MITFMDTKEK SKEVESCLDP QLWHACAGGI VQMPAVNSKV YYFPQGHAEH ACGPVNFRTC 60 PKVPPFVPCR VTAVKYRADP ETDEVYAKLK LIPLNANDVD YDRDVVGGAE TQDKPASFAK 120 TLTQSDANNG GGFSVPRYCA ETIFPRLDYS VDPPVQNILA KDVHGETWKF RHIYRGTPRR 180 HLLTTGWSTF VNHKKLVAGD SIVFLRAENG DLCVGIRRAK KGICGGLETS SGWNPAGGNC 240 HIPYGGFSPF FREDDNRISR NGNSNGLNPS VSMMGKGKVR PEAVSEASNL AANKKPFEVV 300 YYPRASTPEF CVKASLVEAA LQIRWCSGIR FKMAFETEDS SRISWFMGTI SSAQVADPLN 360 WPNSPWRLLQ VTWDEPDLLQ NVRRVSPWLV ELVSNMPAIH FSPFSPPRKK LRLPQQPDFP 420 LDGQIPLSTF PSNLLGPSNT NQFGCLLEST PAGMQGARHA HYGLSLSDLH LSKLQSGLFS 480 TGFPSLDHAA TPMRVSNSIT LQKPNLSENV SCLLTMANST QSSKKLDVGK TPSLVLFGQK 540 ILTEQQISPS SSGDTLSPVL TRNCSSDGNV DKVTNFSDGS GSALHQEGLR EHSSCERFQW 600 CKDNHQETEA GLEIGHCKVF MESEDVGRTM DLSLLRSYDE LHRKLADMFG IEKSEMLSHV 660 LYRDSTGAVK RISDESFSDF TRTAKRLTIL MDSGSNNVGV * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldu_A | 1e-77 | 8 | 391 | 41 | 388 | Auxin response factor 5 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Gma.44757 | 0.0 | hypocotyl| leaf| stem | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, culms, leaves and young panicles. {ECO:0000269|PubMed:17408882}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00461 | DAP | Transfer from AT4G30080 | Download |
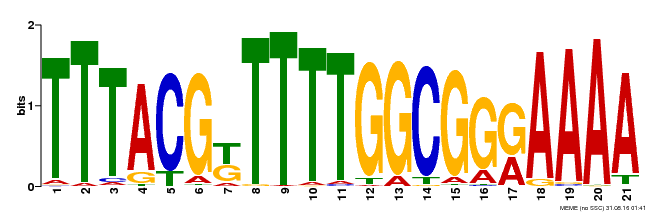
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Glyma.10G053500.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003537296.1 | 0.0 | auxin response factor 18 isoform X1 | ||||
| Refseq | XP_006588784.1 | 0.0 | auxin response factor 18 isoform X1 | ||||
| Swissprot | Q653H7 | 0.0 | ARFR_ORYSJ; Auxin response factor 18 | ||||
| TrEMBL | K7LHL4 | 0.0 | K7LHL4_SOYBN; Auxin response factor | ||||
| STRING | GLYMA10G06080.2 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF637 | 34 | 138 | Representative plant | OGRP526 | 15 | 78 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G30080.1 | 0.0 | auxin response factor 16 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Glyma.10G053500.1.p |
| Entrez Gene | 100796522 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



