 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Glyma.09G265200.1.p | ||||||||
| Common Name | Glyma09g40130.1, LOC100783614 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 821aa MW: 89611.7 Da PI: 6.3731 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 66.1 | 4.7e-21 | 119 | 174 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+++l+L++rqVk+WFqNrR+++k
Glyma.09G265200.1.p 119 KKRYHRHTPQQIQELESLFKECPHPDEKQRLELSRRLNLETRQVKFWFQNRRTQMK 174
688999***********************************************999 PP
| |||||||
| 2 | START | 208.4 | 2.8e-65 | 336 | 560 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S.... CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla.... 77
ela++a++elvk+a+ +ep+W +s e++n de+++++++ + + +ea+r +g+v+ ++ lve+l+d++ +W+e+++
Glyma.09G265200.1.p 336 ELALAAMDELVKMAQTDEPLWIRSLeggrEILNHDEYTRTITPCIGlrpngFVTEASRQTGMVIINSLALVETLMDSN-RWSEMFPcmia 424
5899*************************************999989***9***************************.*********** PP
EEEEEEEECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTC CS
START 78 kaetlevissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksng 160
+ +t evis+g galqlm aelq+lsplvp R++ f+R+++q+ +g w++vdvS+d ++ + + +v +++lpSg+++++++ng
Glyma.09G265200.1.p 425 RTSTAEVISNGingtrnGALQLMHAELQVLSPLVPvREVNFLRFCKQHAEGLWAVVDVSIDTIRDTSGAPTFVNCRRLPSGCVVQDMPNG 514
*****************************************************************999********************** PP
EEEEEEEE-EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 161 hskvtwvehvdlkgrlphwllrslvksglaegaktwvatlqrqcek 206
+skvtwveh++++++++h+l+r+l++sg+ +ga++wvatlqrqce+
Glyma.09G265200.1.p 515 YSKVTWVEHAEYDESQIHQLYRPLLSSGMGFGAQRWVATLQRQCEC 560
********************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.88E-21 | 102 | 176 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 8.9E-22 | 109 | 176 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.783 | 116 | 176 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 2.4E-19 | 117 | 180 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.51E-19 | 118 | 176 | No hit | No description |
| Pfam | PF00046 | 1.1E-18 | 119 | 174 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 151 | 174 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 45.48 | 327 | 563 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 2.2E-35 | 329 | 560 | No hit | No description |
| CDD | cd08875 | 6.82E-126 | 331 | 559 | No hit | No description |
| SMART | SM00234 | 5.6E-48 | 336 | 560 | IPR002913 | START domain |
| Pfam | PF01852 | 6.8E-57 | 336 | 560 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.81E-24 | 588 | 813 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 821 aa Download sequence Send to blast |
MSFGGFLETK QSGGGGGRIV ADIPYSNNSN NIMPSSAISQ PRLATPTLVK SMFNSPGLSL 60 ALQSDIDGKR DVNRLMPENF EQNGLRRNRE EEHESRSGSD NMDGGSGDDF DAADNPPRKK 120 RYHRHTPQQI QELESLFKEC PHPDEKQRLE LSRRLNLETR QVKFWFQNRR TQMKTQLERH 180 ENSLLRQEND KLRAENMSMR EAMRNPICTN CGGPAMIGEI SLEEQHLRIE NARLKDELDR 240 VCALAGKFLG RPISSLTGSI GPPLPNSSLE LGVGSNGFGG LSTVPSTMPD FGVGISSPLA 300 MVSPSSTRPT TTATTTLVTP PSGFDNRSIE RSIVLELALA AMDELVKMAQ TDEPLWIRSL 360 EGGREILNHD EYTRTITPCI GLRPNGFVTE ASRQTGMVII NSLALVETLM DSNRWSEMFP 420 CMIARTSTAE VISNGINGTR NGALQLMHAE LQVLSPLVPV REVNFLRFCK QHAEGLWAVV 480 DVSIDTIRDT SGAPTFVNCR RLPSGCVVQD MPNGYSKVTW VEHAEYDESQ IHQLYRPLLS 540 SGMGFGAQRW VATLQRQCEC LAILISSAVP SREHSAISSG GRRSMLKLAQ RMTNNFCAGV 600 CASTVHKWNK LNAGNVGEDV RVMTRKSVDD PGEPPGIVLS AATSVWLPVS PQRLFDFLRD 660 ERLRSEWDIL SNGGPMQEMA HIAKGQDHAN CVSLLRASAI NANQSSMLIL QETCTDASGS 720 LVVYAPVDIP AMHVVMNGGD SAYVALLPSG FAIVPDGSVE ENGGASQQRA ASGGCLLTVA 780 FQILVNSLPT AKLTVESVET VNNLISCTVQ KIKSALHCES * |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, leaves and floral buds. {ECO:0000269|PubMed:10402424}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
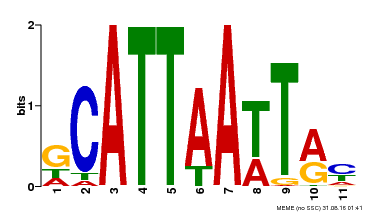
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Glyma.09G265200.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KT031166 | 0.0 | KT031166.1 Glycine max clone HN_CCL_121 homeodomain/HOMEOBOX transcription factor (Glyma09g40130.1) mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003534596.1 | 0.0 | homeobox-leucine zipper protein ANTHOCYANINLESS 2 isoform X1 | ||||
| Refseq | XP_006587870.1 | 0.0 | homeobox-leucine zipper protein ANTHOCYANINLESS 2 isoform X1 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A0K2CTQ2 | 0.0 | A0A0K2CTQ2_SOYBN; Homeodomain/HOMEOBOX transcription factor (Fragment) | ||||
| TrEMBL | I1L6R2 | 0.0 | I1L6R2_SOYBN; Uncharacterized protein | ||||
| STRING | GLYMA09G40130.1 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1192 | 34 | 106 | Representative plant | OGRP145 | 15 | 136 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Glyma.09G265200.1.p |
| Entrez Gene | 100783614 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



