 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Glyma.08G303900.2.p | ||||||||
| Common Name | GLYMA_08G303900, LOC100800166 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 526aa MW: 57805.8 Da PI: 6.8769 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 55 | 1.5e-17 | 331 | 377 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
hn ErrRRdriN+++ L++l+P++ K +Ka++Le+A+eY+ksLq
Glyma.08G303900.2.p 331 VHNLSERRRRDRINEKMKALQQLIPHS-----SKTDKASMLEEAIEYLKSLQ 377
6*************************8.....6******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.280.10 | 3.5E-19 | 324 | 384 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 6.41E-20 | 325 | 388 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 18.47 | 327 | 376 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.09E-15 | 330 | 375 | No hit | No description |
| Pfam | PF00010 | 4.7E-15 | 331 | 377 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 2.1E-18 | 333 | 382 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0009693 | Biological Process | ethylene biosynthetic process | ||||
| GO:0010244 | Biological Process | response to low fluence blue light stimulus by blue low-fluence system | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0010928 | Biological Process | regulation of auxin mediated signaling pathway | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 526 aa Download sequence Send to blast |
MNNSVPDWNF GSDSCVTSTN QKKPMMGVDQ ELVELLWQNG QVVHSQTHRK PVVNSSNLIQ 60 DDETVSWIQY PLEDPLEQEF CSNLLSELAP CEVESYKQIK PFEEGKFTKL DASSAPHVTV 120 SSQLPTMKSS SFQDFSGIPI PAPRFHVSDS PQKNNNHLGG SCKVQNFSHF SAPLNVSSAS 180 ANVHFGDKIT GNMSRNEIRE CSLMTVGSSY CGSNHIPQDP DASRASSNGV WTTTLSVEPE 240 AVRDDVPKTI PQKEKGKSEM LEPAVTSSSG GSGSSLGKTC SLSTRNQGQK RKGIDVEESE 300 EQSEDTELKS ALGNKSSQRT GSARRNRAAE VHNLSERRRR DRINEKMKAL QQLIPHSSKT 360 DKASMLEEAI EYLKSLQLQL QLMWMGSGMA PIMFPGIQHY MSQMGMGMAR PPFPPPIHNP 420 MQLPRVPLDK SVSASQTPNQ TLMCQNPILG AFNYQNQMQN PALSEQYVRY MGYHLMQNAS 480 QPMNVFRYGP QGVQHSQTMI TPSSNSSGNM SGAANIDEAV SGKMG* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 335 | 340 | ERRRRD |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Gma.13444 | 0.0 | cotyledon| leaf | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00082 | ChIP-seq | Transfer from AT3G59060 | Download |
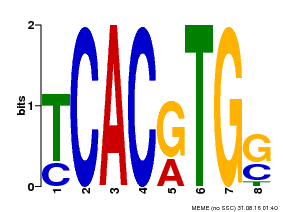
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Glyma.08G303900.2.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC235243 | 0.0 | AC235243.1 Glycine max strain Williams 82 clone GM_WBb0032L23, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003532070.2 | 0.0 | transcription factor PIF4 isoform X1 | ||||
| Refseq | XP_006586042.1 | 0.0 | transcription factor PIF4 isoform X1 | ||||
| Refseq | XP_028245818.1 | 0.0 | transcription factor PIF4-like isoform X1 | ||||
| TrEMBL | A0A445JM89 | 0.0 | A0A445JM89_GLYSO; Transcription factor PHYTOCHROME INTERACTING FACTOR-LIKE 13 isoform B | ||||
| TrEMBL | I1KXW5 | 0.0 | I1KXW5_SOYBN; Uncharacterized protein | ||||
| STRING | GLYMA08G41620.1 | 0.0 | (Glycine max) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G43010.2 | 9e-46 | phytochrome interacting factor 4 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Glyma.08G303900.2.p |
| Entrez Gene | 100800166 |



