 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Glyma.08G221800.3.p | ||||||||
| Common Name | GLYMA_08G221800 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 264aa MW: 29115.2 Da PI: 6.7489 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 48.8 | 9.7e-16 | 226 | 260 | 1 | 33 |
GATA 1 CsnCgttk..TplWRrgpdgnktLCnaCGlyyrkk 33
C++Cg ++ Tp++Rrgp+g++tLCnaCGl+++ k
Glyma.08G221800.3.p 226 CRHCGISEksTPMMRRGPEGPRTLCNACGLMWANK 260
*****99999**********************876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00979 | 2.8E-11 | 82 | 117 | IPR010399 | Tify domain |
| PROSITE profile | PS51320 | 14.532 | 82 | 117 | IPR010399 | Tify domain |
| Pfam | PF06200 | 1.3E-12 | 87 | 116 | IPR010399 | Tify domain |
| PROSITE profile | PS51017 | 12.927 | 150 | 192 | IPR010402 | CCT domain |
| Pfam | PF06203 | 2.3E-15 | 150 | 192 | IPR010402 | CCT domain |
| SMART | SM00401 | 3.1E-5 | 220 | 263 | IPR000679 | Zinc finger, GATA-type |
| SuperFamily | SSF57716 | 2.04E-10 | 221 | 260 | No hit | No description |
| Gene3D | G3DSA:3.30.50.10 | 5.8E-14 | 224 | 260 | IPR013088 | Zinc finger, NHR/GATA-type |
| CDD | cd00202 | 2.34E-11 | 225 | 260 | No hit | No description |
| Pfam | PF00320 | 1.2E-13 | 226 | 260 | IPR000679 | Zinc finger, GATA-type |
| PROSITE pattern | PS00344 | 0 | 226 | 253 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0030154 | Biological Process | cell differentiation | ||||
| GO:0045944 | Biological Process | positive regulation of transcription from RNA polymerase II promoter | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005667 | Cellular Component | transcription factor complex | ||||
| GO:0000977 | Molecular Function | RNA polymerase II regulatory region sequence-specific DNA binding | ||||
| GO:0001085 | Molecular Function | RNA polymerase II transcription factor binding | ||||
| GO:0001228 | Molecular Function | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding | ||||
| GO:0003682 | Molecular Function | chromatin binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 264 aa Download sequence Send to blast |
MDGIHGGDSR IHITDGQHPI HVPYVQEHEH HGLHHISNGN GIDDDHNDGG DTNCGGSESM 60 EGEVPSNHGN LPDNHAVMMD QGGDSGDQLT LSFQGQVYVF DSVSPEKVQA VLLLLGGREI 120 PPTMPAMPVS PNHNNRGYTG TPQKFSVPQR LASLIRFREK RKERNYDKKI RYTVRKEVAL 180 RMQRNKGQFT SSKSNNDESA SNATNWGMDE NWTADNSGSQ QQDIVCRHCG ISEKSTPMMR 240 RGPEGPRTLC NACGLMWANK SFV* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Gma.14008 | 0.0 | cotyledon| hypocotyl| leaf | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Predominantly expressed in shoot apices, inflorescences and roots. {ECO:0000269|PubMed:14966217}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that specifically binds 5'-GATA-3' or 5'-GAT-3' motifs within gene promoters. {ECO:0000250, ECO:0000269|PubMed:14966217}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00198 | DAP | Transfer from AT1G51600 | Download |
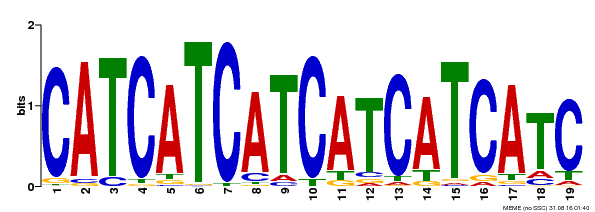
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Glyma.08G221800.3.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015036 | 1e-125 | AP015036.1 Vigna angularis var. angularis DNA, chromosome 3, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_014634655.1 | 0.0 | GATA transcription factor 28 isoform X3 | ||||
| Refseq | XP_028244718.1 | 0.0 | GATA transcription factor 28-like isoform X3 | ||||
| Swissprot | Q8H1G0 | 1e-123 | GAT28_ARATH; GATA transcription factor 28 | ||||
| TrEMBL | A0A0R0IQT6 | 0.0 | A0A0R0IQT6_SOYBN; Uncharacterized protein | ||||
| TrEMBL | A0A445JIE6 | 0.0 | A0A445JIE6_GLYSO; GATA transcription factor 28 isoform C | ||||
| STRING | GLYMA08G23720.2 | 0.0 | (Glycine max) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G51600.2 | 1e-117 | ZIM-LIKE 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Glyma.08G221800.3.p |



