 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Glyma.08G201900.2.p | ||||||||
| Common Name | GLYMA_08G201900 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 827aa MW: 91180.2 Da PI: 6.4002 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 57.7 | 2e-18 | 5 | 63 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++ +++ps +r++L +++ +++ +q+kvWFqNrR +ek+
Glyma.08G201900.2.p 5 KYVRYTPEQVEALERLYHDCPKPSSIRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 63
5679*****************************************************97 PP
| |||||||
| 2 | START | 181.6 | 4.6e-57 | 149 | 357 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT..E CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg..g 89
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s++++g a+ra+g+v +++ v+e+l+d++ W ++++++++l+v+ ++ g
Glyma.08G201900.2.p 149 IAEETLAEFLSKATGTAVEWVQMPGMKPGPDSIGIVAISHGCTGVAARACGLVGLEPT-RVAEILKDRPLWFRDCRAVDVLNVLPTAngG 237
7899******************************************************.8888888888****************9999* PP
EEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SS CS
START 90 alqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvdlkgr 175
+++l +++l+a+++l+p Rdf+ +Ry+ l++g++vi+++S+ ++q+ p+ +++vRae+lpSg+li+p+++g+s +++v+h+dl+ +
Glyma.08G201900.2.p 238 TIELLYMQLYAPTTLAPaRDFWLLRYTSVLEDGSLVICERSLKNTQNGPSmppVQHFVRAEMLPSGYLIRPCEGGGSIIHIVDHMDLEPW 327
************************************************9988899*********************************** PP
XXHHHHHHHHHHHHHHHHHHHHHHTXXXXX CS
START 176 lphwllrslvksglaegaktwvatlqrqce 205
+++++lr+l++s+++ ++kt++a+l+++++
Glyma.08G201900.2.p 328 SVPEVLRPLYESSTVLAQKTTMAALRHLRQ 357
*************************99876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.613 | 1 | 64 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.1E-15 | 2 | 68 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.28E-16 | 5 | 65 | No hit | No description |
| SuperFamily | SSF46689 | 2.35E-16 | 5 | 66 | IPR009057 | Homeodomain-like |
| Pfam | PF00046 | 5.7E-16 | 6 | 63 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 3.4E-18 | 7 | 63 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 2.33E-6 | 57 | 96 | No hit | No description |
| PROSITE profile | PS50848 | 25.069 | 139 | 353 | IPR002913 | START domain |
| CDD | cd08875 | 3.39E-81 | 143 | 358 | No hit | No description |
| SMART | SM00234 | 9.3E-43 | 148 | 358 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.02E-38 | 148 | 358 | No hit | No description |
| Gene3D | G3DSA:3.30.530.20 | 2.9E-23 | 148 | 354 | IPR023393 | START-like domain |
| Pfam | PF01852 | 1.2E-54 | 149 | 357 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 2.31E-5 | 385 | 479 | No hit | No description |
| SuperFamily | SSF55961 | 2.31E-5 | 509 | 584 | No hit | No description |
| Pfam | PF08670 | 3.3E-50 | 683 | 825 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 827 aa Download sequence Send to blast |
MDNGKYVRYT PEQVEALERL YHDCPKPSSI RRQQLIRECP ILSNIEPKQI KVWFQNRRCR 60 EKQRKESSRL QAVNRKLTAM NKLLMEENDR LQKQVSQLVY ENGYFRQHTQ ITTQATKDTN 120 CESVVTSGQH NLTTQHPPRD ASPAGLLSIA EETLAEFLSK ATGTAVEWVQ MPGMKPGPDS 180 IGIVAISHGC TGVAARACGL VGLEPTRVAE ILKDRPLWFR DCRAVDVLNV LPTANGGTIE 240 LLYMQLYAPT TLAPARDFWL LRYTSVLEDG SLVICERSLK NTQNGPSMPP VQHFVRAEML 300 PSGYLIRPCE GGGSIIHIVD HMDLEPWSVP EVLRPLYESS TVLAQKTTMA ALRHLRQISH 360 EVSQSNVTGW GRRPAALRAL SQRLSRGFNE ALNGFTDEGW TTISNDGVDD VTILVNSSPD 420 KLMGLNLSFA NGFPSVSNAV LCAKASMLLQ NVPPAILLRF LREHRSEWAD NNMDAYTAAA 480 IKVGPCSLSG SCVGNFGGQV ILPLAHTIEH EEFLEVIKLE GIAHSPEDTI MPREMFLLQL 540 CSGMDENAVG TCAELISAPI DASFADDAPL LPSGFRIIPL ESGKEASSPN RTLDLASSLD 600 VGPSGNRASN GSAGNSSCMR SVMTIAFEFA FESHMQEHVT SMARQYVRSI ISSVQRVALA 660 LSPSHLSSHA GLRSPLGTPE AQTLAHWICN SYRCYLGVEL LKSNNEGNES LLKSLWHHSD 720 AILCCTLKAL PVFTFSNQAG LDMLETTLVA LQDITLEKIF DDHGRKILFS EFPQIIQQGF 780 ACLQGGICLS SMGRPVSYER VVAWKVLNEE ENAHCICFMF VNWSFV* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Gma.7462 | 0.0 | hypocotyl| meristem| seed coat | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Highly expressed the developing vascular elements and the adaxial portion of cotyledons. Expressed in developing ovules, stamens and carpels. Expressed in procambium and shoot meristem. {ECO:0000269|PubMed:14701930, ECO:0000269|PubMed:15705957}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
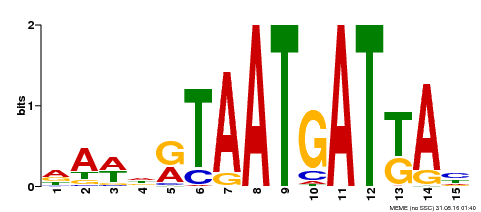
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Glyma.08G201900.2.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003531652.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 | ||||
| Refseq | XP_028244497.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | A0A0R0IPU8 | 0.0 | A0A0R0IPU8_SOYBN; Uncharacterized protein | ||||
| TrEMBL | A0A445JHH8 | 0.0 | A0A445JHH8_GLYSO; Homeobox-leucine zipper protein ATHB-15 isoform B | ||||
| STRING | GLYMA08G21610.1 | 0.0 | (Glycine max) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.1 | 0.0 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Glyma.08G201900.2.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



