 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Glyma.08G029400.1.p | ||||||||
| Common Name | GLYMA_08G029400, Glyma08g03330.1, MYB127 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 268aa MW: 29387 Da PI: 9.7759 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 45 | 2.5e-14 | 98 | 142 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE+ +++ + ++lG+g+W+ I+r + ++Rt+ q+ s+ qky
Glyma.08G029400.1.p 98 PWTEEEHRIFLVGLEKLGKGDWRGISRNFVTTRTPTQVASHAQKY 142
8*******************************************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.60.10 | 6.0E-4 | 3 | 20 | IPR001878 | Zinc finger, CCHC-type |
| PROSITE profile | PS51294 | 18.527 | 91 | 147 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 9.33E-18 | 92 | 147 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 6.2E-18 | 94 | 145 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 1.2E-11 | 95 | 141 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 3.5E-11 | 95 | 145 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 9.7E-12 | 98 | 142 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 6.62E-10 | 98 | 143 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006338 | Biological Process | chromatin remodeling | ||||
| GO:0006357 | Biological Process | regulation of transcription from RNA polymerase II promoter | ||||
| GO:0035066 | Biological Process | positive regulation of histone acetylation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003682 | Molecular Function | chromatin binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0003713 | Molecular Function | transcription coactivator activity | ||||
| GO:0004402 | Molecular Function | histone acetyltransferase activity | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 268 aa Download sequence Send to blast |
MGRKCSHCGT IGHNSRTCTS LRGATTSFVG LRLFGVQLDS TNCVSIKKSF SMDSLPSSSS 60 SSFSSSRLTI DENSDRTSFG YLSDGLLARA QERKKGVPWT EEEHRIFLVG LEKLGKGDWR 120 GISRNFVTTR TPTQVASHAQ KYFLRLATID KKKRRSSLFD LVGSNKAGSN SVSAHQKDES 180 KCEVKNNDAA TLSLLGRITY FQQENKSSEY KQETFDNCSY QSQPGEEHQV AVLNLDLTLA 240 APKAKTLQEQ NKSSPSAPFL LGPISVT* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 150 | 154 | KKKRR |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Gma.22805 | 0.0 | cotyledon| hypocotyl| leaf| root | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in all tissues, with the highest level in senescent leaves. {ECO:0000269|PubMed:12172034}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription repressor that binds to 5'-TATCCA-3' elements in gene promoters. Contributes to the sugar-repressed transcription of promoters containing SRS or 5'-TATCCA-3' elements. Transcription repressor involved in a cold stress response pathway that confers cold tolerance. Suppresses the DREB1-dependent signaling pathway under prolonged cold stress. DREB1 responds quickly and transiently while MYBS3 responds slowly to cold stress. They may act sequentially and complementarily for adaptation to short- and long-term cold stress (PubMed:20130099). {ECO:0000269|PubMed:12172034, ECO:0000269|PubMed:20130099}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00565 | DAP | Transfer from AT5G56840 | Download |
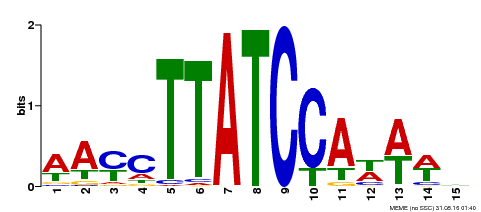
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Glyma.08G029400.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Repressed by sucrose and gibberellic acid (GA) (PubMed:12172034). Induced by cold stress in roots and shoots. Induced by salt stress in shoots. Down-regulated by abscisic aci (ABA) in shoots (PubMed:20130099). {ECO:0000269|PubMed:12172034, ECO:0000269|PubMed:20130099}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KT031194 | 0.0 | KT031194.1 Glycine max clone HN_CCL_90 MYB/HD-like transcription factor (Glyma08g03330.1) mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001235940.1 | 0.0 | MYB transcription factor MYB127 | ||||
| Refseq | XP_028244981.1 | 0.0 | transcription factor MYBS3-like | ||||
| Swissprot | Q7XC57 | 3e-49 | MYBS3_ORYSJ; Transcription factor MYBS3 | ||||
| TrEMBL | A0A445J9B8 | 0.0 | A0A445J9B8_GLYSO; Transcription factor MYBS3 | ||||
| TrEMBL | Q0PJI8 | 0.0 | Q0PJI8_SOYBN; MYB transcription factor MYB127 | ||||
| STRING | GLYMA08G03330.1 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3213 | 32 | 71 | Representative plant | OGRP394 | 16 | 99 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G56840.1 | 2e-54 | MYB_related family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Glyma.08G029400.1.p |
| Entrez Gene | 778052 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



