 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Glyma.07G080300.1.p | ||||||||
| Common Name | GLYMA_07G080300, LOC100804760 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | TCP | ||||||||
| Protein Properties | Length: 336aa MW: 35580.1 Da PI: 10.0398 | ||||||||
| Description | TCP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCP | 119.8 | 3.3e-37 | 65 | 166 | 2 | 104 |
TCP 2 agkkdrhskihTkvggRdRRvRlsaecaarfFdLqdeLGfdkdsktieWLlqqakpaikeltgtssssasec.eaesssssasnsssgka 90
+ k+ +++++hTkv+gR+RR+R++a+caar+F+L++eLG+++d++ti WLl+ a+pai+++tgt++ +a ++ +++ + + s +++
Glyma.07G080300.1.p 65 PQKRASTKDRHTKVEGRGRRIRMPATCAARIFQLTRELGHKSDGETIRWLLEHAEPAIIAATGTGTVPAIAMsVNGT--LKIPTTSPSDQ 152
57999***********************************************************7777744421111..11111111112 PP
TCP 91 aksaakskksqksa 104
++ +++++k++k++
Glyma.07G080300.1.p 153 EPGDQPERKKRKRP 166
22222111111111 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03634 | 1.9E-31 | 70 | 173 | IPR005333 | Transcription factor, TCP |
| PROSITE profile | PS51369 | 27.141 | 71 | 125 | IPR017887 | Transcription factor TCP subgroup |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008361 | Biological Process | regulation of cell size | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 336 aa Download sequence Send to blast |
MADEASGDVD LRLGEPKAPK EEPETEEGSA IRVMPVAVHV PAGMSMSMSK ALAQAQAQAQ 60 AQAQPQKRAS TKDRHTKVEG RGRRIRMPAT CAARIFQLTR ELGHKSDGET IRWLLEHAEP 120 AIIAATGTGT VPAIAMSVNG TLKIPTTSPS DQEPGDQPER KKRKRPANSA YVDINDAAVS 180 ISTGLASSNN NNQNAATTTT MTIPQHTAIP LPQGMVPVWA IPSNAVVPAA GAFFVVPQTA 240 SFQHQPQFFT IARPISAFVS SLVTPVQPQP TSITSSSNAN ASSTAPSSKS AARATVMAPT 300 TTTATTTTTQ MLRDFSLEIY DKQELQFMSR ASSKH* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 158 | 165 | ERKKRKRP |
| 2 | 159 | 164 | RKKRKR |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Gma.4458 | 0.0 | cotyledon| root | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in archespores, pollen mother cell, ovule primordia and megaspore mother cells. {ECO:0000269|PubMed:25527103}. | |||||
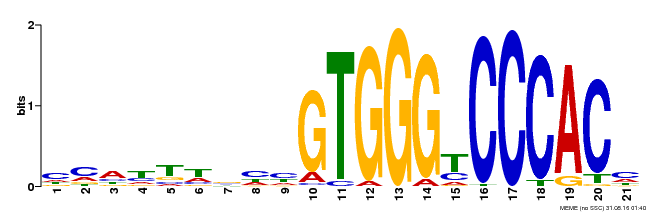
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00316 | DAP | Transfer from AT2G45680 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Glyma.07G080300.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC235048 | 0.0 | AC235048.1 Glycine max strain Williams 82 clone GM_WBb0045O22, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003529971.1 | 0.0 | transcription factor TCP9 | ||||
| Refseq | XP_028239692.1 | 0.0 | transcription factor TCP9-like | ||||
| Swissprot | O64647 | 8e-94 | TCP9_ARATH; Transcription factor TCP9 | ||||
| TrEMBL | A0A445JTY0 | 0.0 | A0A445JTY0_GLYSO; Transcription factor TCP9 | ||||
| TrEMBL | K7L0D3 | 0.0 | K7L0D3_SOYBN; Uncharacterized protein | ||||
| STRING | GLYMA07G08710.2 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2118 | 30 | 86 | Representative plant | OGRP3178 | 11 | 29 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G45680.1 | 7e-73 | TCP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Glyma.07G080300.1.p |
| Entrez Gene | 100804760 |



