 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Glyma.06G049200.1.p | ||||||||
| Common Name | GLYMA_06G049200, LOC100802454 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | AP2 | ||||||||
| Protein Properties | Length: 661aa MW: 73129.3 Da PI: 7.7572 | ||||||||
| Description | AP2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 50.9 | 3.7e-16 | 297 | 356 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrd.pse.ng..kr.krfslgkfgtaeeAakaaiaarkkleg 55
s+y+GV++++++gr++A+++d ++ g ++ ++++lg ++ +e+Aa+a++ a++k++g
Glyma.06G049200.1.p 297 SQYRGVTRHRWTGRYEAHLWDnSCKkEGqtRKgRQVYLGGYDMEEKAARAYDLAALKYWG 356
78*******************988886678446*************************98 PP
| |||||||
| 2 | AP2 | 47.2 | 5.6e-15 | 399 | 450 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s y+GV+++++ grW A+I + +k +lg+f t eeAa+a++ a+ k++g
Glyma.06G049200.1.p 399 SIYRGVTRHHQHGRWQARIGRVAG---NKDLYLGTFSTQEEAAEAYDIAAIKFRG 450
57****************988532...5*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 2.68E-16 | 297 | 366 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 4.17E-21 | 297 | 363 | No hit | No description |
| Pfam | PF00847 | 9.0E-13 | 297 | 356 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 3.1E-15 | 298 | 364 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 1.1E-27 | 298 | 370 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 18.782 | 298 | 364 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 2.0E-6 | 299 | 310 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 5.82E-26 | 399 | 460 | No hit | No description |
| SuperFamily | SSF54171 | 9.15E-18 | 399 | 460 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 8.4E-10 | 399 | 450 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.032 | 400 | 458 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 1.4E-18 | 400 | 458 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 4.9E-34 | 400 | 464 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 2.0E-6 | 440 | 460 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007276 | Biological Process | gamete generation | ||||
| GO:0010492 | Biological Process | maintenance of shoot apical meristem identity | ||||
| GO:0042127 | Biological Process | regulation of cell proliferation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 661 aa Download sequence Send to blast |
MKRMNESNNT DDGNNHNWLG FSLSPHMKME VTSAATVSDN NVPTTFYMSP SHMSNSGMCY 60 SVGENGNFHS PLTVMPLKSD GSLGILEALN RSQTQVMVPT SSPKLEDFLG GATMGTHEYG 120 NHERGLSLDS IYYNSQNAEA QPNRNLLSHP FRQQGHVNVE THPYYSVFAC RGLYQAPSEE 180 EATKETHVSV MPQMTGGGLQ NWVAPTREYS THQQILEQQM NCGIWNERSG VSVGTVGCGE 240 LQSLSLSMSP GSQSSCVTAP SGTDSVAVDA KKRGHAKLGQ KQPVHRKSID TFGQRTSQYR 300 GVTRHRWTGR YEAHLWDNSC KKEGQTRKGR QVYLGGYDME EKAARAYDLA ALKYWGPSTH 360 INFSIENYQV QLEEMKNMSR QEYVAHLRRK SSGFSRGASI YRGVTRHHQH GRWQARIGRV 420 AGNKDLYLGT FSTQEEAAEA YDIAAIKFRG ANAVTNFDIS RYDVERIMAS SNLLAGELAR 480 RNKDNDPRNE AIDYNKSVVT SVNNGETVQV QARNNNENDS EWKMVLFNHP LQQQANGSDH 540 KIMNCGNSRN SAFSMALQDL IGVDSVGSEQ HNMLDDSSKI GTHFSNPSSL VTSLSSSREA 600 SPEKMGPSLL FPKPPPMETK IVNPIGTSVT SWLPSPTVQM RPSPAISLSH LPVFAAWTDT 660 * |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expressed in floral primordia, in STM-negative region, then in sepal primordia. As sepal develops, progressively confined to a basal core before disappearing. Present in stamen primordia, then confined to a central region as they become stalked and develop locules. Later reduced to procambial cells as stamen mature. From petal primordia, expressed on the lateral edges of developing petals and finally confined to petal epidermis before disappearing. Present in carpel primordia, then in inner side of carpels especially in the placenta. Strong levels in ovules primordia and young ovules, then localized in integuments initiation zone before being confined to inner integument cells that will differentiate into the endothelium. Expressed in the distal half of the funiculus throughout ovule development and later extends into the chalaza. After fertilization, expression shift to the embryo. First on the apical part at the globular stage, then in cotyledons primordia, and later in cotyledons during the torpedo stage. As cotyledons grow out, expression becomes limited to a plane separating adaxial and abaxial parts. Excluded from the embryonic central region (ECR). In seedlings, found in leaf primordia then in central and lateral actively developing regions of extending leaves. {ECO:0000269|PubMed:10656774, ECO:0000269|PubMed:8742707, ECO:0000269|PubMed:9671577}. | |||||
| Uniprot | TISSUE SPECIFICITY: Mostly expressed in developing flowers. Also present in mature flowers, siliques and seedlings, but not in mature roots, leaves and stems. Expressed in ovules and in vegetative and floral primordia. {ECO:0000269|PubMed:15988559, ECO:0000269|PubMed:8742706, ECO:0000269|PubMed:8742707}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CAC[AG]N[AT]TNCCNANG-3'. Required for the initiation and growth of ovules integumenta, and for the development of female gametophyte. Plays a critical role in the development of gynoecium marginal tissues (e.g. stigma, style and septa), and in the fusion of carpels and of medial ridges leading to ovule primordia. Also involved in organs initiation and development, including floral organs. Maintains the meristematic competence of cells and consequently sustains expression of cell cycle regulators during organogenesis, thus controlling the final size of each organ by controlling their cell number. Regulates INO autoinduction and expression pattern. As ANT promotes petal cell identity and mediates down-regulation of AG in flower whorl 2, it functions as a class A homeotic gene. {ECO:0000269|PubMed:10528263, ECO:0000269|PubMed:10639184, ECO:0000269|PubMed:10948255, ECO:0000269|PubMed:11041883, ECO:0000269|PubMed:12183381, ECO:0000269|PubMed:12271029, ECO:0000269|PubMed:12655002, ECO:0000269|PubMed:8742706, ECO:0000269|PubMed:8742707, ECO:0000269|PubMed:9001406, ECO:0000269|PubMed:9093862, ECO:0000269|PubMed:9118807}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00088 | SELEX | Transfer from AT4G37750 | Download |
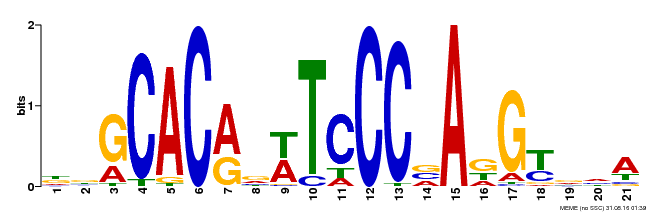
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Glyma.06G049200.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006581282.1 | 0.0 | AP2-like ethylene-responsive transcription factor ANT | ||||
| Refseq | XP_028235004.1 | 0.0 | AP2-like ethylene-responsive transcription factor ANT | ||||
| Swissprot | Q38914 | 1e-160 | ANT_ARATH; AP2-like ethylene-responsive transcription factor ANT | ||||
| TrEMBL | A0A445K4W6 | 0.0 | A0A445K4W6_GLYSO; AP2-like ethylene-responsive transcription factor ANT isoform A | ||||
| TrEMBL | K7KT57 | 0.0 | K7KT57_SOYBN; Uncharacterized protein | ||||
| STRING | GLYMA06G05170.2 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF916 | 34 | 114 | Representative plant | OGRP112 | 17 | 209 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G37750.1 | 1e-150 | AP2 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Glyma.06G049200.1.p |
| Entrez Gene | 100802454 |



