 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Glyma.05G200800.4.p | ||||||||
| Common Name | GLYMA_05G200800, LOC100815956 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 859aa MW: 94845.1 Da PI: 6.8191 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 75.3 | 7.1e-24 | 160 | 261 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFk 83
f+k+lt sd++++g +++ +++a+e+ ++ + +++l+ +d++g++W++++i+r++++r++l++GW+ Fv++++L +gD ++F
Glyma.05G200800.4.p 160 FCKTLTASDTSTHGGFSVLRRHADEClppldmtKQ--PPTQELVAKDLHGNEWRFRHIFRGQPRRHLLQSGWSVFVSSKRLVAGDAFIFL 247
99****************************97433..34459************************************************ PP
E-SSSEE..EEEEE-S CS
B3 84 ldgrsefelvvkvfrk 99
+ +++el+v+v+r+
Glyma.05G200800.4.p 248 --RGENGELRVGVRRA 261
..449999*****996 PP
| |||||||
| 2 | Auxin_resp | 116.8 | 1.9e-38 | 286 | 368 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
a+ha+ t+++F+v+Y+Pr+s++eF+v++++++++lk+++++GmRfkm+fe+e+++e+r++Gt+vg++d+d+ rWp+SkWrsLk
Glyma.05G200800.4.p 286 AWHAILTGTMFTVYYKPRTSPAEFIVPYDQYMESLKNNYTIGMRFKMRFEGEEAPEQRFTGTIVGIEDADTKRWPKSKWRSLK 368
79*******************************************************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 1.6E-41 | 156 | 275 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 2.09E-40 | 156 | 289 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 7.87E-22 | 158 | 260 | No hit | No description |
| PROSITE profile | PS50863 | 11.998 | 160 | 262 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 9.1E-22 | 160 | 261 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 2.1E-23 | 160 | 262 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 4.0E-36 | 286 | 368 | IPR010525 | Auxin response factor |
| PROSITE profile | PS51745 | 22.873 | 734 | 816 | IPR000270 | PB1 domain |
| SuperFamily | SSF54277 | 1.28E-10 | 736 | 811 | No hit | No description |
| Pfam | PF02309 | 2.2E-8 | 780 | 822 | IPR033389 | AUX/IAA domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010227 | Biological Process | floral organ abscission | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 859 aa Download sequence Send to blast |
MATSEVSIKG NSVNGKGDNS SGGYTNDVRN GSGGGEARNS SSSSSSARDA EAALYRELWH 60 ACAGPLVTVP REGERVFYFP QGHIEQVEAS TNQVAEQHMP VYDLPPKILC RVINVMLKAE 120 PDTDEVFAQV TLLPEPNQDE NAVEKEGPPA APPRFHVHSF CKTLTASDTS THGGFSVLRR 180 HADECLPPLD MTKQPPTQEL VAKDLHGNEW RFRHIFRGQP RRHLLQSGWS VFVSSKRLVA 240 GDAFIFLRGE NGELRVGVRR AMRQQGNVPS SVISSHSMHL GVLATAWHAI LTGTMFTVYY 300 KPRTSPAEFI VPYDQYMESL KNNYTIGMRF KMRFEGEEAP EQRFTGTIVG IEDADTKRWP 360 KSKWRSLKVR WDETSNIPRP ERVSQWKIEP ALAPPALNPL PMPRPKRPRS NVVPSSPDSS 420 VLTREASSKV SVDPLPTSGF QRVLQGQELS TLRGNFAESN ESDTVEKSAV WPPVADDEKI 480 DVSTSRRYGS DSWMSMGRHE LTYPDLLSGF GTHGDHSSHP SFVDQNGPVA NVGRKHLLDC 540 EGKHNVLSPW SGVPSSLSLN LLDSNTKGSA QGGDTTYQVR GNLRYSSAFG EYPMLHGHKV 600 EHSHGNFLMP PPPSTPYESP RSRELLPKPI SGKPCEVSKP KDSDCKLFGI SLLSSPIAPE 660 PSVSQRNVPS EPVGHMHTTS HQQRAFDNDQ KSEHSRGGSK PADGLLIDDH EKVLQTSQTH 720 LKDIQAKSHS GSARSCTKVH KKGIALGRSV DLTKFSDYGE LIAELDQLFE FGGLLTSPQK 780 DWLIVYTDNE GDMMLVGDDP WQEFVAMVRK IYIYPKEEIQ KMSPGTLSSK NEENQSASEG 840 ATDTQEIKCQ LNNSASDT* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 1e-178 | 53 | 390 | 18 | 354 | Auxin response factor 1 |
| 4ldw_A | 1e-178 | 53 | 390 | 18 | 354 | Auxin response factor 1 |
| 4ldw_B | 1e-178 | 53 | 390 | 18 | 354 | Auxin response factor 1 |
| 4ldx_A | 1e-178 | 53 | 390 | 18 | 354 | Auxin response factor 1 |
| 4ldx_B | 1e-178 | 53 | 390 | 18 | 354 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Gma.2563 | 0.0 | cotyledon| epicotyl| flower| hypocotyl| leaf| meristem| pod| root| somatic embryo| stem | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expression increases during floral development. Expressed during ovary development with the highest expression on the third day after the flower fully opened. {ECO:0000269|PubMed:21735233}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in root, leaf and stem (PubMed:26716451, PubMed:21735233). Also expressed in flower and fruit (PubMed:26716451). Expressed in flower buds about three days before opening including stamen, petal and sepal with the highest in ovary (PubMed:21735233). {ECO:0000269|PubMed:21735233, ECO:0000269|PubMed:26716451}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that binds specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs) (By similarity). Could act as transcriptional activator or repressor. Involved in the control of fruit ripening process. Regulates expression of a number of ripening regulators, transcription factors, and ethylene biosynthesis and signaling components (PubMed:26716451, PubMed:26959229). May act as a transcriptional repressor of auxin-responsive genes (PubMed:26716451). {ECO:0000255|RuleBase:RU004561, ECO:0000269|PubMed:26716451, ECO:0000269|PubMed:26959229}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00574 | DAP | Transfer from AT5G62000 | Download |
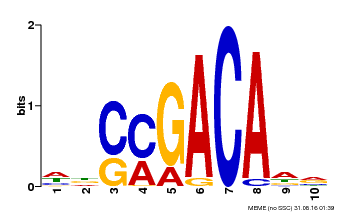
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Glyma.05G200800.4.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By tomato fruit ripening. Expression is up-regulated by plant hormone auxin in tomato fruits. {ECO:0000269|PubMed:26716451}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC235431 | 0.0 | AC235431.1 Glycine max strain Williams 82 clone GM_WBb0144O21, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003525433.1 | 0.0 | auxin response factor 2B-like isoform X1 | ||||
| Refseq | XP_028233438.1 | 0.0 | auxin response factor 2B-like isoform X1 | ||||
| Swissprot | K4DF01 | 0.0 | ARF2B_SOLLC; Auxin response factor 2B | ||||
| TrEMBL | A0A0B2QCB4 | 0.0 | A0A0B2QCB4_GLYSO; Auxin response factor | ||||
| TrEMBL | I1K6T1 | 0.0 | I1K6T1_SOYBN; Auxin response factor | ||||
| STRING | GLYMA05G38540.4 | 0.0 | (Glycine max) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62000.3 | 0.0 | auxin response factor 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Glyma.05G200800.4.p |
| Entrez Gene | 100815956 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



