 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Glyma.03G225200.1.p | ||||||||
| Common Name | GLYMA_03G225200 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 419aa MW: 46765.1 Da PI: 7.003 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 54.5 | 2.7e-17 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g W++eEde+l++ ++++G g+W+++++ g+ R++k+c++rw +yl
Glyma.03G225200.1.p 14 KGLWSPEEDEKLLRHITKYGHGCWSSVPKQAGLQRCGKSCRLRWINYL 61
678*******************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 46.6 | 7.8e-15 | 67 | 110 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
rg +++eE+ l++++++ lG++ W+ Ia+ ++ gRt++++k+ w++
Glyma.03G225200.1.p 67 RGTFSQEEENLIIELHAVLGNR-WSQIAAQLP-GRTDNEIKNLWNS 110
899*******************.*********.***********97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 4.2E-28 | 6 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 18.926 | 9 | 61 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.61E-29 | 12 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.7E-13 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.0E-15 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.03E-11 | 17 | 61 | No hit | No description |
| PROSITE profile | PS51294 | 25.948 | 62 | 116 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 7.6E-26 | 65 | 116 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.8E-13 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.0E-13 | 67 | 111 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.74E-9 | 69 | 109 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 419 aa Download sequence Send to blast |
MGRHSCCYKQ KLRKGLWSPE EDEKLLRHIT KYGHGCWSSV PKQAGLQRCG KSCRLRWINY 60 LRPDLKRGTF SQEEENLIIE LHAVLGNRWS QIAAQLPGRT DNEIKNLWNS CLKKKLRQRG 120 IDPVTHKPLS EVENGEEGKT RSQELSNELN LLNSESFKSD EGSYEQRASS IAPKAYEMEG 180 SCSSKINNTK NDTNLMSNCS NKDMFLDSYT TSCQPSDLMG NYPLHITDTL PTNSNSCHWF 240 SQTARPFDIN SNSEFTITSN VMSILPPTTT SFLPSTSFCY KPSLGVPSED ISAPSFGLNG 300 PSYWEASASN NSNGSNNTSD GSRELTTTSS KNSVLSSWEL TEEAKWSEYL HNPMLMLAAP 360 ESLCNQIRPA THLIPDNTLG AIILPDSKNQ QQQQQQQSQN SCIFSKDVQK LTAAFGHI* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 9e-29 | 12 | 116 | 5 | 108 | B-MYB |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 111 | 117 | LKKKLRQ |
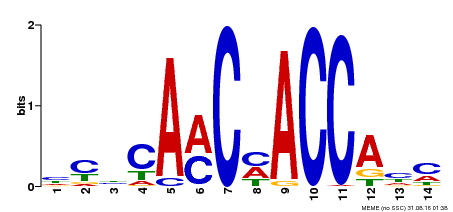
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00427 | DAP | Transfer from AT4G01680 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Glyma.03G225200.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KT031185 | 0.0 | KT031185.1 Glycine max clone HN_CCL_13 MYB/HD-like transcription factor (Glyma03g38410.1) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001304642.2 | 0.0 | MYB/HD-like transcription factor | ||||
| TrEMBL | I1JQY1 | 0.0 | I1JQY1_SOYBN; Uncharacterized protein | ||||
| STRING | GLYMA03G38410.1 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1083 | 33 | 103 | Representative plant | OGRP5 | 17 | 1784 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G01680.3 | 6e-94 | myb domain protein 55 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Glyma.03G225200.1.p |



