 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Gh_D10G0079 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 759aa MW: 83275.9 Da PI: 6.8956 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 34.9 | 3.5e-11 | 712 | 757 | 4 | 47 |
S-HHHHHHHHHHHHHTTTT-HHHHHHHHT...TTS-HHHHHHHHHHH CS
Myb_DNA-binding 4 WTteEdellvdavkqlGggtWktIartmg...kgRtlkqcksrwqky 47
WT eE++ l+++ +++G+ Wk+I + k+Rt ++k++w+++
Gh_D10G0079 712 WTHEEEQALIKGYREYGTQ-WKLILESYRdilKKRTQVDLKDKWRNL 757
*****************99.*************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 14.223 | 704 | 759 | IPR017930 | Myb domain |
| SMART | SM00717 | 0.0016 | 708 | 759 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 5.07E-11 | 711 | 757 | IPR009057 | Homeodomain-like |
| CDD | cd11660 | 2.63E-16 | 712 | 759 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 3.5E-10 | 712 | 757 | IPR009057 | Homeodomain-like |
| Pfam | PF13921 | 4.7E-9 | 712 | 758 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009901 | Biological Process | anther dehiscence | ||||
| GO:0010152 | Biological Process | pollen maturation | ||||
| GO:0043067 | Biological Process | regulation of programmed cell death | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 759 aa Download sequence Send to blast |
MGDSFPLFND RDPAVLSTLK QYLSQPNNND EDNGDGYATF CFHYKGAILR SIEQRTGNST 60 VPDAVLDSLI LIENLDRRRG VLPSESMKAA FCAVAVHCTV SCLPVSWDNY FDAFHRIWGL 120 RIKSLEESGK SDLISSELVQ WGTVIEAGLW DLETSQRLSS INTRGKALLR IKGYLEEAFR 180 SMSPALSRLA SASTTEPTVD HVASANPNPS TDEGNMDNFV SASAQVPSSP HPCIDERNMD 240 NVVYASAQVP SSPHPCSDEG KMDNVVSASA EASPSPHPCA DKGNVDNVIS SSAQVPSSPH 300 PCTNKKKKIR RASFQARHKP RPRCKQPRGV VITDMEEDQP LCTKNGTPSS FEVNGCLQVA 360 SKTRSAALLD GVTDSPSEAL EVLETVALVM AGKNFHPKGS VEDSNKDKGD PTTSFYPTTH 420 HCTEKGKQIL MVNSQTSGNP NSSHRHCEGP AATADIEEDR ALSTPDIDKL RDALTSSIAD 480 LTATVADPLP KALEVAARLV SFMEAAKSVS TDGAEGDVRK EKGVPAAPMN CNFEQAQAER 540 RAPDKAIRED QDKMARSVPA SSVGHSVEPS QAKEGNETFS RPKKVPKRNL MKRNDAAHTH 600 ERRAPYDGIR EDYNKVDGSV PAPTVNHTAE PSHSKEGNEA VIRPKKVRKC SIMERNDTAH 660 TSEWEDSIDG SDGGTSGCSN RCTLPSPKTN HVSPLKESEP QKQKNRRKTN WWTHEEEQAL 720 IKGYREYGTQ WKLILESYRD ILKKRTQVDL KDKWRNLSK |
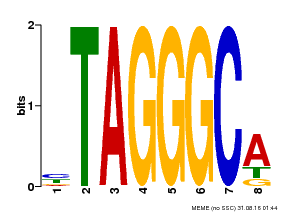
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00052 | PBM | Transfer from AT1G15720 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016717472.1 | 0.0 | PREDICTED: uncharacterized protein LOC107930355 isoform X1 | ||||
| TrEMBL | A0A1U8LSB9 | 0.0 | A0A1U8LSB9_GOSHI; uncharacterized protein LOC107930355 isoform X1 | ||||
| STRING | Gorai.011G008700.1 | 0.0 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM22142 | 4 | 5 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G15720.1 | 3e-13 | TRF-like 5 | ||||



