 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Gh_D08G1195 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 986aa MW: 108589 Da PI: 5.7068 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 90.3 | 1.5e-28 | 580 | 630 | 2 | 52 |
RWP-RK 2 ekeisledlskyFslpikdAAkeLgvclTvLKriCRqyGIkRWPhRkiksl 52
ek+isle+l++yF +++k+AAk+Lgvc+T++KriCRq+GI+RWP+Rki+++
Gh_D08G1195 580 EKSISLEVLQQYFTGSLKNAAKSLGVCPTTMKRICRQHGISRWPSRKINKV 630
799**********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51519 | 17.188 | 565 | 650 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 1.5E-25 | 583 | 630 | IPR003035 | RWP-RK domain |
| SuperFamily | SSF54277 | 9.81E-23 | 884 | 974 | No hit | No description |
| PROSITE profile | PS51745 | 23.282 | 890 | 972 | IPR000270 | PB1 domain |
| Gene3D | G3DSA:3.10.20.240 | 1.3E-25 | 890 | 972 | No hit | No description |
| SMART | SM00666 | 7.5E-25 | 890 | 972 | IPR000270 | PB1 domain |
| Pfam | PF00564 | 1.4E-16 | 891 | 971 | IPR000270 | PB1 domain |
| CDD | cd06407 | 2.45E-37 | 891 | 971 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 986 aa Download sequence Send to blast |
MCEPEEDNAC PFPPKQLQPQ HQGIMDFDDL DLQSSWPFDQ LTFLSNPTSP FVTSSSSEQP 60 CSPLWAFSDD DKLGSAAAAG YNLFVTCTPN PVNENAKEEN DNRGLPSLFL GLLPLENPDS 120 YCVIKERMTR ALRYFKESTE QHVLAQVWAP VKDGGRYVLT TLGQPFVLDP HSSGLHQYRM 180 VSLMYMFSVD GESDVQLGLP GRVFRQKLPE WTPNVQYYSS REYSRLNHAL HYNVQGTLAL 240 PVFEPSGQSC VGVLELIMTS QKINYAPEVD KVCKALEAVN LKSSEILGYP STQICNESRQ 300 NALAEILEIL TVVCETHKLP LAQTWVPCRH RKVLVHGGGL KKSCTSFDGS CMGQVCMSTT 360 DVAFYVVDAH MWGFRDACLE HHLQKGQGVA GRAFLSLNSC FCSDITQFCK TEYPLVHYAR 420 MFKLTSCFAI CLRSAYTRDD DYVLEFFLPP AIADCNEQQA LLGSILTTMK QHFQSLMVAS 480 GAELEEDEGS IEIIGASAEE RFVTRLECIP IPPPVKSPPK NNTSPNKGEL QLDSSKQQLT 540 VNFDPSAHGG RVVASGSHNL DCPLQNKDLK KPERKRGKTE KSISLEVLQQ YFTGSLKNAA 600 KSLGVCPTTM KRICRQHGIS RWPSRKINKV NRSLTKLKHV IESVHGTDGA FGLTSLANSP 660 LPVGVDSISW PTGLNGSNQQ NSPNSRPFEH KGEKNDSPTC LTPQSNGQVV VEDQLLGGRT 720 LSPEPFPQQN GLSSHFDKGV KRSRRGSTSR EESAGTPISY SSCQGSLGIE NAATKASFGS 780 IHDQCLKAHG SPELAFQQLL GEPNISAMFS MPKVLVAAEL EEPIGGMLVE AAGSSKDLRN 840 LCPIADIGVN EQFPESSWTP PPCSDLALKQ AMSTFTQPTP RVTAREETKS VTIKATYRED 900 IIRFRISMSS GILELKEEVA NRLKLEVGAC DIKYLDDDNE WVLIACDADL EECIDVSRSS 960 GNNIIRLCIH DTMANLESSC ESTGEL |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, leaves, flowers and siliques. Detected in root hairs, emerging secondary roots, vascular tissues, leaf parenchyma cells and stomata. {ECO:0000269|PubMed:18826430}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in regulation of nitrate assimilation and in transduction of the nitrate signal. {ECO:0000269|PubMed:18826430}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
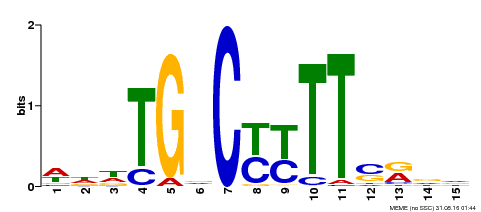
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Not regulated by the N source or by nitrate. {ECO:0000269|PubMed:18826430}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016694696.1 | 0.0 | PREDICTED: protein NLP6-like isoform X1 | ||||
| Swissprot | Q84TH9 | 0.0 | NLP7_ARATH; Protein NLP7 | ||||
| TrEMBL | A0A1U8JWP2 | 0.0 | A0A1U8JWP2_GOSHI; protein NLP6-like isoform X1 | ||||
| STRING | Gorai.004G131700.1 | 0.0 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2507 | 26 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24020.1 | 0.0 | NIN like protein 7 | ||||



