 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Gh_D01G0951 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 946aa MW: 104239 Da PI: 5.0055 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 77.1 | 2e-24 | 150 | 251 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE.. CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefel 92
f+k+lt sd++++g +++p++ ae+ ++++++ ++l+++d++ ++W++++iyr++++r++lt+GW+ Fv +++L++gD+v+F ++++ +l
Gh_D01G0951 150 FCKTLTASDTSTHGGFSVPRRAAEKLfpsldySMQPPT-QELVVRDLHDNTWTFRHIYRGQPKRHLLTTGWSLFVGSKRLRAGDSVLFI--RDEKSQL 244
99*********************999*****9555554.49************************************************..4577778 PP
EEEEE-S CS
B3 93 vvkvfrk 99
+v+v+r+
Gh_D01G0951 245 LVGVRRA 251
*****97 PP
| |||||||
| 2 | Auxin_resp | 113.5 | 1.9e-37 | 276 | 359 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalk.vkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
aahaa+++s+F+++YnPr+++seFv+++ +++k+++ ++vsvGmRf m+fete+s +rr++Gt+vg+sdldp+rWp+SkWr+L+
Gh_D01G0951 276 AAHAAANRSPFTIFYNPRSCPSEFVIPLPRYRKSVYgSQVSVGMRFGMMFETEESGKRRYMGTIVGISDLDPLRWPGSKWRNLQ 359
79**********************************99********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 9.68E-47 | 137 | 279 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 4.0E-42 | 144 | 264 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 4.47E-22 | 149 | 250 | No hit | No description |
| SMART | SM01019 | 4.6E-24 | 150 | 252 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 7.0E-22 | 150 | 251 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.183 | 150 | 252 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 2.9E-32 | 276 | 359 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 2.7E-9 | 825 | 915 | IPR033389 | AUX/IAA domain |
| PROSITE profile | PS51745 | 24.573 | 829 | 913 | IPR000270 | PB1 domain |
| SuperFamily | SSF54277 | 6.21E-9 | 831 | 908 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009908 | Biological Process | flower development | ||||
| GO:0009942 | Biological Process | longitudinal axis specification | ||||
| GO:0010305 | Biological Process | leaf vascular tissue pattern formation | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0048507 | Biological Process | meristem development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016020 | Cellular Component | membrane | ||||
| GO:0042802 | Molecular Function | identical protein binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 946 aa Download sequence Send to blast |
MGSVVEEKIK QGGLVNVGAQ STLLEEMKLL KEMQDQSGTR KAINSELWHA CAGPLVSLPQ 60 VGSLVYYFPQ GHSEQQVAVS TKRMATSQIP NYPNLPSQLM CQVHNVTLHA DRDTDEIYAQ 120 MSLQPVNSEK DVFPIPDFGL KLSKHPNEFF CKTLTASDTS THGGFSVPRR AAEKLFPSLD 180 YSMQPPTQEL VVRDLHDNTW TFRHIYRGQP KRHLLTTGWS LFVGSKRLRA GDSVLFIRDE 240 KSQLLVGVRR ANRQQTTLPS SVLSADSMHI GVLAAAAHAA ANRSPFTIFY NPRSCPSEFV 300 IPLPRYRKSV YGSQVSVGMR FGMMFETEES GKRRYMGTIV GISDLDPLRW PGSKWRNLQV 360 EWDEPGCNDK QNRVSAWEIE TPESLFIFPS LTSSLKRPLY PGFSAAESEW GSLMKRPLLQ 420 FPENGNGNLP YSMSNLCSEQ LMKMMLKPQL VNHPGIFASP LQQIADVKVP PLEEMKNLQS 480 KSHPKPQVIQ SENMLIENRN LSHPVPDQPD PITSNMSKIN ANGNPHPANI LTQAGTGSSN 540 EKLKLDSKHS AEQLTSTSEC NEEKLVASTV NTTMSNQLSF PTQPHIPLQV QNNPWSIQSQ 600 LDSSVLQAHQ MLVSQADIST LNSFLPFSDT DEWTSNLSSC QPLSGAYKSP GPIPMVGLQD 660 SSAVFPVETD DSLTTVGEEI WDQKLNNCRV SSQADQLASF TEQDPCSLNS GGVRDLSDDS 720 NNQSGIYSSC LNIDVSNGCS TVIDPFVSSA ILDEFCSLKD ADFQNPSDCL VGNFSSCQDV 780 QSQITSASLA DSQAFSRQDL PDSSGGNIDF DDSGLLQNNS WKQTAPRVRT YTKVQKAGSV 840 GRSIDVTSFK NYDELISAIE CMFGLKGLLD DPRGSGWKLV YVDYENDVLL VGDDPWEEFV 900 GCVRCIRILS PTEVQQMSEE GMKLLNSAAT VQGINGSNSE GSNANA |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldu_A | 0.0 | 3 | 384 | 6 | 392 | Auxin response factor 5 |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: In early embryo and during organ development. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in the whole plant with a lower expression in leaves. Detected in embryo axis, provascular tissues, procambium and some differentiated vascular regions of mature organs. {ECO:0000269|PubMed:10476078}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). Seems to act as transcriptional activator. Formation of heterodimers with Aux/IAA proteins may alter their ability to modulate early auxin response genes expression. Mediates embryo axis formation and vascular tissues differentiation. Functionally redundant with ARF7. May be necessary to counteract AMP1 activity. {ECO:0000269|PubMed:12036261, ECO:0000269|PubMed:14973283, ECO:0000269|PubMed:17553903}. | |||||
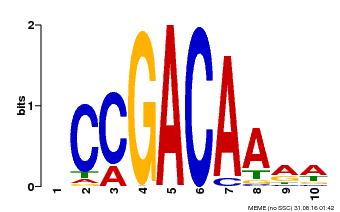
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00153 | DAP | Transfer from AT1G19850 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016707387.1 | 0.0 | PREDICTED: auxin response factor 5-like | ||||
| Swissprot | P93024 | 0.0 | ARFE_ARATH; Auxin response factor 5 | ||||
| TrEMBL | A0A1U8KY74 | 0.0 | A0A1U8KY74_GOSHI; Auxin response factor | ||||
| STRING | Gorai.002G124400.1 | 0.0 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5808 | 27 | 46 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G19850.1 | 0.0 | ARF family protein | ||||



