 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Gh_A12G2645 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 314aa MW: 35203.4 Da PI: 7.5441 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 53.4 | 6e-17 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd + v +++++G+g+W++I+ g+ R+ k+c++rw +yl
Gh_A12G2645 14 KGPWTPEEDIIMVSYIQEHGPGNWRSIPTNTGLLRCSKSCRLRWTNYL 61
79******************************99************97 PP
| |||||||
| 2 | Myb_DNA-binding | 50.5 | 4.9e-16 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg++T++E++ ++++ ++lG++ W++Ia++++ Rt++++k++w+++l
Gh_A12G2645 67 RGNFTEQEEKMIIHLQALLGNR-WAAIASYLP-QRTDNDIKNYWNTHL 112
89********************.*********.************996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 9.2E-24 | 5 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 17.393 | 9 | 61 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.25E-30 | 11 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 8.4E-13 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.4E-15 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.64E-10 | 16 | 61 | No hit | No description |
| PROSITE profile | PS51294 | 24.775 | 62 | 116 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.4E-25 | 65 | 117 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 6.5E-16 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.1E-15 | 67 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.05E-11 | 69 | 112 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0080167 | Biological Process | response to karrikin | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 314 aa Download sequence Send to blast |
MGRPPCCDKI GVKKGPWTPE EDIIMVSYIQ EHGPGNWRSI PTNTGLLRCS KSCRLRWTNY 60 LRPGIKRGNF TEQEEKMIIH LQALLGNRWA AIASYLPQRT DNDIKNYWNT HLKKKLKKVE 120 TGVDGQNQEG FSSAQSVSKG QWERRLQTDI RMAKQALSDA LSLDKPNSLT NSIEFKLSHP 180 YLRPSQSQSS TAYASSAENI SRLLQNWMKN PPKPAPAPRQ TKSVETMTQN SSSSDEGALS 240 EATPEGFDSF LTFNSSNSDN ISHESVSVAE NSVFQDESKP NLRLIEKWLL DDASGQAHDD 300 DLINMSLQDS AALF |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1h8a_C | 2e-26 | 14 | 116 | 27 | 128 | MYB TRANSFORMING PROTEIN |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Ghi.12171 | 0.0 | boll| ovule | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expression in flowers increases as the flowers develop. {ECO:0000269|PubMed:1840903}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in flowers, leaves and weakly in seed pods. {ECO:0000269|PubMed:1840903}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. {ECO:0000305}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00395 | DAP | Transfer from AT3G47600 | Download |
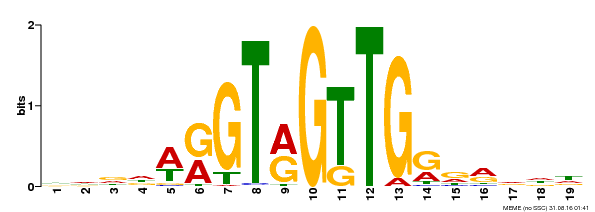
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016722141.1 | 0.0 | PREDICTED: myb-related protein 306-like isoform X1 | ||||
| Swissprot | P81392 | 1e-125 | MYB06_ANTMA; Myb-related protein 306 | ||||
| TrEMBL | A0A1U8M5P5 | 0.0 | A0A1U8M5P5_GOSHI; myb-related protein 306-like isoform X1 | ||||
| STRING | Gorai.008G192900.1 | 0.0 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4 | 28 | 2646 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G47600.1 | 1e-116 | myb domain protein 94 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



