 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Gh_A11G0926 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 812aa MW: 89390.6 Da PI: 7.5602 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 50.4 | 5e-16 | 104 | 148 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT+eE+ ++++a k++G W +I +++g ++t+ q++s+ qk+
Gh_A11G0926 104 RERWTEEEHNRFLEALKLYGRA-WQRIEEHIG-TKTAVQIRSHAQKF 148
78******************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 6.73E-17 | 98 | 154 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 21.091 | 99 | 153 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 9.0E-17 | 102 | 151 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 7.3E-13 | 103 | 151 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.4E-13 | 104 | 147 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.3E-8 | 104 | 144 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.61E-9 | 106 | 149 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 812 aa Download sequence Send to blast |
MTSYLTQKGH VPVAHATLAQ CHPLFHTWWY GYNRGNLAGG EVTRRKRVPR GKCGGAFTDL 60 TSQNGNLWLR FEMNELSSSM AICQGFGVYG FVLTRKPYTI TKQRERWTEE EHNRFLEALK 120 LYGRAWQRIE EHIGTKTAVQ IRSHAQKFFS KLEKEALTKG IPTGQALDIE IPPPRPKRKP 180 RNPYPRKMSA ATSVQMGAKD GKSETPVSSL HCKQAFNLEK EPLPERPSRD EKSSNLKEIL 240 DDNCSEVFTL LHEANCSSMS SVNKNFIPTS TVLRNSCPLR EFVPSLKETI NKDTSKPSNL 300 ENSCTSYEKP AQVQRKDDMD GATCTDEMQA THNNPQHVAV HVLDGSPGTC APNPSMDITF 360 QDSIFHPMGD IHGQVNLFAN PAASATTAHQ NNAPRSTHQA FPTFHTPFMH LQPNQEDYRS 420 FLHVSSTFSS LVVSTLLQNP AAHAAASFAA TFWPYANVQN SDDSPACDQK GFPSRQMNSA 480 TSMAAIAAAT VAAATAWWAA HGLLSVCAPL QTGCTCAPAS TAAVTPMENG EAPAANMERK 540 MYAGQDPSMQ DQRLDPQYAE AMQCQPSASK SPTSSSSDRE ESGEAKANTE VKATAATEPQ 600 DPNKTKNGKQ VDRSSCGSNT PSSSDVEIDV LEKNKEDKED PKPADANHPQ VECSNRRGRS 660 STNLSDSWKE VSEGGRLAFQ ALFSREVLPQ SFSPPHDRKN KEQQKESVGE DEQNSDQNNG 720 ETSILHFSSE AFRSCSDHQG VEKKALSKAK NIVDKSLLTI GLGHEKLKAR QTGFKPYKRC 780 SVEAKESRVM NTGSQGEEKD PKRIRLEGEA PT |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 174 | 180 | RPKRKPR |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in leaves, roots, stems, flowers and siliques. {ECO:0000269|PubMed:19095940, ECO:0000269|PubMed:19218364}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in the circadian clock. Binds to the promoter region of APRR1/TOC1 and TCP21/CHE to repress their transcription. Represses both CCA1 and itself. {ECO:0000269|PubMed:12015970, ECO:0000269|PubMed:19095940, ECO:0000269|PubMed:19218364, ECO:0000269|PubMed:9657154}. | |||||
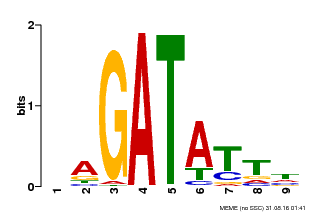
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00654 | PBM | Transfer from LOC_Os08g06110 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Circadian-regulation with peak levels occurring around 1 hour after dawn. Up-regulated by APRR1/TOC1 and transiently by light treatment. Down-regulated by APRR5, APRR7 and APRR9. {ECO:0000269|PubMed:12574129, ECO:0000269|PubMed:19095940, ECO:0000269|PubMed:19218364, ECO:0000269|PubMed:19286557, ECO:0000269|PubMed:20233950, ECO:0000269|PubMed:9657154}. | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HQ526180 | 8e-46 | HQ526180.1 Gossypium herbaceum clone NBRI_E_1400 simple sequence repeat marker, mRNA sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016709652.1 | 0.0 | PREDICTED: protein LHY-like isoform X1 | ||||
| Swissprot | Q6R0H1 | 1e-137 | LHY_ARATH; Protein LHY | ||||
| TrEMBL | A0A1U8L4V6 | 0.0 | A0A1U8L4V6_GOSHI; protein LHY-like isoform X1 | ||||
| STRING | Gorai.007G113900.1 | 0.0 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5576 | 26 | 45 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01060.3 | 1e-112 | MYB_related family protein | ||||



