 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Gh_A09G0075 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 374aa MW: 41992.4 Da PI: 5.6223 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 53.3 | 4.9e-17 | 215 | 261 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
hn ErrRRdriN+++ L+el+P++ K +Ka++L +A+eY+ksLq
Gh_A09G0075 215 VHNLSERRRRDRINEKMRALQELIPHC-----NKTDKASMLDEAIEYLKSLQ 261
6*************************8.....6******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF47459 | 1.44E-21 | 209 | 277 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 18.457 | 211 | 260 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 2.09E-18 | 214 | 265 | No hit | No description |
| Pfam | PF00010 | 2.4E-14 | 215 | 261 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 2.1E-21 | 215 | 271 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 7.2E-19 | 217 | 266 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0009693 | Biological Process | ethylene biosynthetic process | ||||
| GO:0010244 | Biological Process | response to low fluence blue light stimulus by blue low-fluence system | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0010928 | Biological Process | regulation of auxin mediated signaling pathway | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 374 aa Download sequence Send to blast |
MNHCSSDWNF DSDLPISNQK KLLGQDNELV ELLWQNGKVV LQSQTHKKQD DETVSWIQNP 60 FEESFEKEMF SNFFSEFPAY DPMDHQHEQQ EDDKSLKHKN NLMQREVNEL SGMAIGSSHC 120 GSNQVRNDGD LSRGSSNGIG TTSTGLSVGT SKDDEDDYNG GQVENEKGTS VGVTSSQKRK 180 NGDGREDYEC QSEFAEDKPV RRSGSYRRRS RAAEVHNLSE RRRRDRINEK MRALQELIPH 240 CNKTDKASML DEAIEYLKSL QSQLQVMWMG NGMAPMMFPG IQHYMSPMAM GTAPPTMPSI 300 QNQRFIQNPT FSEQYARFLG FQHMQTASQP INMFGYGSQT TPQSPTVLGF SNGSNQLNGG 360 ITAANNTSLS GKIG |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 206 | 223 | RRRSRAAEVHNLSERRRR |
| 2 | 219 | 224 | ERRRRD |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00082 | ChIP-seq | Transfer from AT3G59060 | Download |
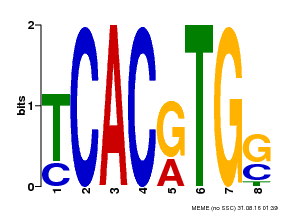
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HQ525802 | 1e-76 | HQ525802.1 Gossypium herbaceum clone NBRI_D_EYT27PB02DXEGO simple sequence repeat marker, mRNA sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016668077.1 | 0.0 | PREDICTED: transcription factor PIF4-like isoform X2 | ||||
| Refseq | XP_016668078.1 | 0.0 | PREDICTED: transcription factor PIF4-like isoform X3 | ||||
| TrEMBL | A0A1U8HPW9 | 0.0 | A0A1U8HPW9_GOSHI; transcription factor PIF4-like isoform X2 | ||||
| TrEMBL | A0A1U8HVV3 | 0.0 | A0A1U8HVV3_GOSHI; transcription factor PIF4-like isoform X3 | ||||
| STRING | Gorai.006G008800.1 | 0.0 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3468 | 25 | 61 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G43010.2 | 4e-53 | phytochrome interacting factor 4 | ||||



