 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Gh_A05G2443 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 556aa MW: 60453.6 Da PI: 4.7989 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 55.2 | 1.7e-17 | 40 | 87 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT Ed +l+d+vk++G g+W+++ ++ g+ R++k+c++rw ++l
Gh_A05G2443 40 KGPWTSAEDAILIDYVKKHGEGNWNAVQKHSGLFRCGKSCRLRWANHL 87
79******************************************9986 PP
| |||||||
| 2 | Myb_DNA-binding | 53.9 | 4.1e-17 | 93 | 136 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g++T+eE++l++++++++G++ W++ a++++ gRt++++k++w++
Gh_A05G2443 93 KGAFTQEEEQLIIELHAKMGNK-WARMAAHLP-GRTDNEIKNYWNT 136
799*******************.*********.***********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 16.037 | 35 | 87 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.55E-30 | 39 | 134 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.4E-15 | 39 | 89 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.4E-15 | 40 | 87 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.2E-24 | 41 | 94 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 6.41E-12 | 42 | 87 | No hit | No description |
| PROSITE profile | PS51294 | 26.872 | 88 | 142 | IPR017930 | Myb domain |
| SMART | SM00717 | 2.3E-17 | 92 | 140 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.4E-16 | 93 | 136 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.11E-12 | 95 | 136 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 2.8E-26 | 95 | 141 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 556 aa Download sequence Send to blast |
MSHTKHERED GMLSKDQTES PYIDDGSCGG GGAGGVVLKK GPWTSAEDAI LIDYVKKHGE 60 GNWNAVQKHS GLFRCGKSCR LRWANHLRPN LKKGAFTQEE EQLIIELHAK MGNKWARMAA 120 HLPGRTDNEI KNYWNTRIKR RQRAGLPLYP PEVSLQALQE SHSTSVVNGL DKGPNDILQN 180 NSYEIPDVIF DSLKANQNVL PYVPELPVLS ASSMLMKGLG SSQYCGFMQP TIHRQKRLRE 240 SPAFFPGYTG AVKNECPLFM QFQDDISDKA AGSFGLSFPI EPDPATKNSQ PFGVFPGSHA 300 LSNGNFSASE PPLEAVKLEL PSLQYPETEL GNWGTLTCPP PLLESVDAFI QSPPPTSGLE 360 SDSLSPRNSG LLDALLHEAK TLSSAKNHAS DKSSNSSTPG DIAEGSNFNI CETEWEKCGE 420 PLSPMGNSAT SLFSECISAS GSSLDEQPPA ETVTESHLKS EPADCVLTPE IQKEAPIRLD 480 SSRPDTLLAS NWLEQGSGYD KDQAILTDAI SSLLGDDLRS EYKNMAEGTS ISSQAWGLDS 540 CAWNNMPAVC QMSELP |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 6e-32 | 38 | 141 | 5 | 107 | B-MYB |
| Search in ModeBase | ||||||
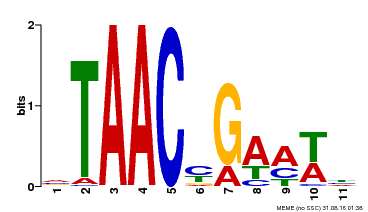
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AJ459146 | 1e-68 | AJ459146.1 Gossypium hirsutum partial myb118 gene, clone pGhMYB118. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016748196.1 | 0.0 | PREDICTED: transcription factor GAMYB-like isoform X1 | ||||
| Refseq | XP_016748197.1 | 0.0 | PREDICTED: transcription factor GAMYB-like isoform X1 | ||||
| Refseq | XP_016748198.1 | 0.0 | PREDICTED: transcription factor GAMYB-like isoform X1 | ||||
| Refseq | XP_016748199.1 | 0.0 | PREDICTED: transcription factor GAMYB-like isoform X1 | ||||
| Refseq | XP_016748200.1 | 0.0 | PREDICTED: transcription factor GAMYB-like isoform X1 | ||||
| TrEMBL | A0A2P5WEH5 | 0.0 | A0A2P5WEH5_GOSBA; Uncharacterized protein | ||||
| STRING | Gorai.009G301100.1 | 0.0 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3802 | 28 | 59 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.3 | 1e-145 | myb domain protein 33 | ||||



