 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EPS72831.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Lentibulariaceae; Genlisea
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 797aa MW: 91650 Da PI: 8.1597 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 88.1 | 1.3e-27 | 51 | 138 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtraetrtgCkaklkvkkekdgkwevtkleleHnHelap 91
+fY+eYAk++GF++++++s++sk+++e+++++f Cs++g++ e +++ ++r+ +++t+Cka+++vk+++dgkw+++++ +eHnHel p
EPS72831.1 51 SFYQEYAKSMGFTTSIKNSRRSKKTKEFIDAKFACSRYGTTPEYENS---SSRRPGVKKTDCKASMHVKRKRDGKWYIHEFIKEHNHELLP 138
5***************************************9998777...7788889*******************************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 4.4E-25 | 51 | 138 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 1.2E-20 | 259 | 350 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.965 | 538 | 574 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 6.4E-7 | 548 | 573 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 1.3E-8 | 549 | 576 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010018 | Biological Process | far-red light signaling pathway | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 797 aa Download sequence Send to blast |
DGIALNLCDT VKNWDCPMSI CSKGDASGTE DNRSEADDGS IEFESHEVAY SFYQEYAKSM 60 GFTTSIKNSR RSKKTKEFID AKFACSRYGT TPEYENSSSR RPGVKKTDCK ASMHVKRKRD 120 GKWYIHEFIK EHNHELLPAL AYHFHIHRNV KLAEKNNIDI LHAVSERTRK MYVEMCRQSG 180 GIYHPFVCKN KFDSRIEGSR CVALDEGDAR IVMDHFMQLK KENPHFFYAV ELNEEQRVRN 240 LFWIDARNRR DYVSFNDAVF FDTAYIKSNE KIPISLFLGV NHHFQPMLLG CGLLADETKS 300 TFVWLLKTWL GAVGGKSPNV IISGQDKQLT SAIEEIFPSS RHCFPLWHVL ERMPETLGNV 360 LRQHSNFMKK FNKCIFKSLP DEEFDMRWWK MVTRFELQEN EWVQSLYVDR KKWVPAFMSN 420 VFLAGLSPHH RSESLNSVFD KYVHKKLNLK EFLRQYSTML RNRYEEEELA DFDTCHKQAA 480 LKSPSPWEKQ MSTIYTHAIF RKFQVEVLGV VGCHSKKEEE DGVTAAFKVD DYEKGENFAV 540 TYNKAKLEVS CSCLMFEYKG FLCRHSMSVL QRCGLSSIPS SYILKRWTKD AKAKQGAAAT 600 VEETDRVQGR VQRYNDLCRL AIRLGEEGSF CDKNYNIACR ALVEALKNCV EVNNKTAAVD 660 SSNYSSGLRF SEQESLLLHG TSKTNKKNTN KKRKSQQEPP EAVLLEAQDN LEPMGNLSSQ 720 DMTLTGYYAS QQNVQGLLNL MEPPQDVYYV GQQTMPGMLN SIPSRHHDSF YGGAAHHQCL 780 PPGGHLEFRQ TAFPYGP |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 238 | 249 | RNLFWIDARNRR |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
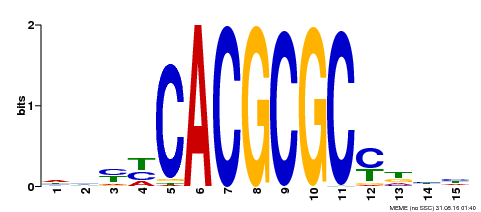
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00434 | DAP | Transfer from AT4G15090 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. {ECO:0000269|PubMed:18033885}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011085456.1 | 0.0 | protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| Refseq | XP_011085457.1 | 0.0 | protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| Swissprot | Q9SWG3 | 0.0 | FAR1_ARATH; Protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| TrEMBL | S8D629 | 0.0 | S8D629_9LAMI; Uncharacterized protein (Fragment) | ||||
| STRING | Migut.H01263.1.p | 0.0 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA496 | 21 | 103 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G15090.1 | 0.0 | FAR1 family protein | ||||



