 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EPS72360.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Lentibulariaceae; Genlisea
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 843aa MW: 91930.2 Da PI: 6.0389 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 66.1 | 4.7e-21 | 123 | 178 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+k+l L++rqVk+WFqNrR+++k
EPS72360.1 123 KKRYHRHTPQQIQELEALFKECPHPDEKQRLELSKRLCLETRQVKFWFQNRRTQMK 178
688999***********************************************999 PP
| |||||||
| 2 | START | 208.9 | 1.9e-65 | 325 | 547 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEEEEEC CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla....kaetlevis 86
ela++a++elvk+++ ++p+W +s e++n++e++++f++ + + +ea+r+sgvv+ ++ lve+l+d++ +W e ++ + +t++vis
EPS72360.1 325 ELALAAMDELVKMVETDDPLWIRSLengrEILNEEEYMRNFTPCIGmkpdgFITEASRESGVVIINSLALVETLMDSN-KWAEIFPcivaRTTTTDVIS 422
5899**************************************99989*******************************.******************** PP
TT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXH CS
START 87 sg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksnghskvtwvehvdlkgrlph 178
g g l+lm ae+q+lsplvp Rd+ f+R+++ql +g w++vdvSvd ++ p +++ R+++lpSg+++++++ng+skvtwveh++++++l+h
EPS72360.1 423 GGmggtrsGSLHLMHAEFQVLSPLVPvRDVHFLRFCKQLAEGAWAVVDVSVDAVRETP--AGYSRCRRLPSGCVVQDTPNGYSKVTWVEHTEYDDSLVH 519
********************************************************99..6************************************** PP
HHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 179 wllrslvksglaegaktwvatlqrqcek 206
+l+r+l++sg +ga +wvatlqrqce+
EPS72360.1 520 QLYRPLINSGFGFGAPRWVATLQRQCEC 547
**************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.28E-20 | 105 | 180 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-21 | 111 | 180 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.216 | 120 | 180 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 9.9E-18 | 121 | 184 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.39E-18 | 122 | 180 | No hit | No description |
| Pfam | PF00046 | 1.4E-18 | 123 | 178 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 155 | 178 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 47.073 | 316 | 550 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 3.85E-37 | 316 | 547 | No hit | No description |
| CDD | cd08875 | 2.93E-111 | 320 | 546 | No hit | No description |
| SMART | SM00234 | 5.0E-56 | 325 | 547 | IPR002913 | START domain |
| Pfam | PF01852 | 1.0E-57 | 325 | 547 | IPR002913 | START domain |
| Gene3D | G3DSA:3.30.530.20 | 5.5E-5 | 430 | 541 | IPR023393 | START-like domain |
| SuperFamily | SSF55961 | 4.4E-24 | 574 | 790 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 843 aa Download sequence Send to blast |
MSFGDNYNIS GGGSARIAPD LDLSYANGAA MPTTGAAIPH PHLLPHSINK PIFNSSATGL 60 SLALQSQSGM ERQGDVGRIG ENYDGNNVGV RNREEENESR SMSDNFDGGS GDDQDAADKP 120 PRKKRYHRHT PQQIQELEAL FKECPHPDEK QRLELSKRLC LETRQVKFWF QNRRTQMKTQ 180 LERHENSLLR QENDKLRSEN LSMRDAMRNP ICTNCGGPAI MGEISLEEQH LRIDNARLKD 240 ELERVCVLAG KFLGRPASSF PVPSMPNSSL ELGVGNNGFG GLNPSMLPLG FPDFVVGGGG 300 GGLPVMPASK TTMNSAPIER SLYLELALAA MDELVKMVET DDPLWIRSLE NGREILNEEE 360 YMRNFTPCIG MKPDGFITEA SRESGVVIIN SLALVETLMD SNKWAEIFPC IVARTTTTDV 420 ISGGMGGTRS GSLHLMHAEF QVLSPLVPVR DVHFLRFCKQ LAEGAWAVVD VSVDAVRETP 480 AGYSRCRRLP SGCVVQDTPN GYSKVTWVEH TEYDDSLVHQ LYRPLINSGF GFGAPRWVAT 540 LQRQCECLAI LMSSSSPAKD HTAISTGGMK SMLKLAQRMT NNFCAGVCAS TVHKWNKLRT 600 ENVDDDVRVM TRKSVDDPGE PQGIVLSAAT SVWLPVTAQR LFDFLRNEHL RSEWDILSNG 660 GPMQEMAHIA KGQDHGNCVS LLRASAMNAN QSNMLILQET CVDAAGSLVV YAPVDIPAMH 720 VVMNGGDSAY VALLPSGFAI VPDGRGGVED GSLLTVAFQI LVNSLPTAKL TVESVETVNN 780 LISCTVQKIK AALRCDEETS SVSSGESRTN LRRKEILATW SELGRGLRPR CKGGGSGIDS 840 GAS |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
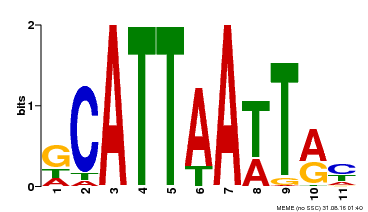
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011093166.1 | 0.0 | homeobox-leucine zipper protein ANTHOCYANINLESS 2 isoform X1 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | S8CZA1 | 0.0 | S8CZA1_9LAMI; Uncharacterized protein | ||||
| STRING | cassava4.1_001764m | 0.0 | (Manihot esculenta) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA1626 | 24 | 65 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G61150.1 | 0.0 | homeodomain GLABROUS 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



