 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EPS60201.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Lentibulariaceae; Genlisea
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 282aa MW: 29567.5 Da PI: 6.9698 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 26.6 | 1.4e-08 | 11 | 57 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT...-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGgg...tWktIartmgkgRtlkqcksrwqky 47
WT e d+++ +a+ + + +W+++a+++ g+++ ++++++ +
EPS60201.1 11 VWTREQDKQFERAIVTYSDDcpdRWEKVASEVA-GKSAGEVRDHYEVL 57
7*****************99*************.***********876 PP
| |||||||
| 2 | Myb_DNA-binding | 42.1 | 2e-13 | 116 | 160 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT++E+ l++ + ++G+g+W++I+r + +Rt+ q+ s+ qky
EPS60201.1 116 AWTEDEHRLFLLGLDKYGKGDWRSISRNFVVTRTPTQVASHAQKY 160
6*******************************************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51293 | 9.086 | 7 | 60 | IPR017884 | SANT domain |
| SMART | SM00717 | 1.3E-7 | 8 | 60 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.8E-5 | 9 | 59 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 2.62E-12 | 10 | 66 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 5.2E-7 | 11 | 57 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.63E-8 | 12 | 58 | No hit | No description |
| PROSITE profile | PS51294 | 17.581 | 108 | 165 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.2E-16 | 111 | 166 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.4E-17 | 113 | 163 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 3.2E-10 | 113 | 163 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-10 | 116 | 159 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 7.32E-10 | 116 | 161 | No hit | No description |
| Pfam | PF00249 | 2.5E-11 | 116 | 160 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 282 aa Download sequence Send to blast |
MPAEESSSSS VWTREQDKQF ERAIVTYSDD CPDRWEKVAS EVAGKSAGEV RDHYEVLLDD 60 VSRIESGFVE LPSYNSSSDG STSHAAEDGK KSNSGESNQG SGKSSRSDQE RRKGIAWTED 120 EHRLFLLGLD KYGKGDWRSI SRNFVVTRTP TQVASHAQKY FIRLNSMNKD RRRSSIHDIT 180 SVNVNAAGAG GGGGGGGGGD VPITGTAAAA SPGAKSNSGK HPPPPGIYGQ PTIGQPVGGP 240 LVSAVGTPVN LPPPGHMAAY GIPGGVVPGA PLTMPPATYH TP |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cjj_A | 1e-15 | 12 | 77 | 11 | 76 | RADIALIS |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that coordinates abscisic acid (ABA) biosynthesis and signaling-related genes via binding to the specific promoter motif 5'-(A/T)AACCAT-3'. Represses ABA-mediated salt (e.g. NaCl and KCl) stress tolerance. Regulates leaf shape and promotes vegetative growth. {ECO:0000269|PubMed:26243618}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00500 | DAP | Transfer from AT5G08520 | Download |
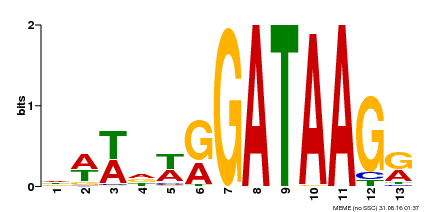
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by salicylic acid (SA) and gibberellic acid (GA) (PubMed:16463103). Triggered by dehydration and salt stress (PubMed:26243618). {ECO:0000269|PubMed:16463103, ECO:0000269|PubMed:26243618}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011070471.1 | 1e-127 | transcription factor SRM1 | ||||
| Refseq | XP_011070472.1 | 1e-127 | transcription factor SRM1 | ||||
| Refseq | XP_020547442.1 | 1e-127 | transcription factor SRM1 | ||||
| Refseq | XP_020547443.1 | 1e-127 | transcription factor SRM1 | ||||
| Swissprot | Q9FNN6 | 1e-100 | SRM1_ARATH; Transcription factor SRM1 | ||||
| TrEMBL | S8C715 | 0.0 | S8C715_9LAMI; Uncharacterized protein (Fragment) | ||||
| STRING | XP_008360003.1 | 1e-125 | (Malus domestica) | ||||
| STRING | XP_008366961.1 | 1e-125 | (Malus domestica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA4149 | 24 | 45 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G08520.1 | 1e-102 | MYB family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



