 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EPS57450.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Lentibulariaceae; Genlisea
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 767aa MW: 88063.7 Da PI: 7.0145 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 82.3 | 7.9e-26 | 91 | 193 | 2 | 91 |
FAR1 2 fYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkk.............tekerrtraetrtgCkaklkvkkekdgkwevtkleleHnH 87
+Y+eYA+++GF++ +++s++sk+++e+++++f Cs++g+++e +k ++++ +ra ++t+Cka+++vk+++dgkw ++++e++HnH
EPS57450.1 91 YYQEYARSTGFNTAIQNSRRSKTSREFIDAKFACSRYGTKREYEKFlnrprsrqggnldPDNASGRRACSKTDCKASMHVKRRSDGKWIIHRFEKDHNH 189
8****************************************9999899999999999887778889********************************* PP
FAR1 88 elap 91
el p
EPS57450.1 190 ELLP 193
*975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 8.6E-24 | 91 | 193 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 4.0E-27 | 291 | 382 | IPR018289 | MULE transposase domain |
| Pfam | PF04434 | 6.0E-5 | 571 | 604 | IPR007527 | Zinc finger, SWIM-type |
| PROSITE profile | PS50966 | 9.396 | 571 | 607 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 5.0E-7 | 582 | 609 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 767 aa Download sequence Send to blast |
MDIDLRLPSG EHDKEIEEEP NIIDGIMVGE DKPINADGVD VSMEIVEAKL LIEAAENEDA 60 SSLHEMDFKE ATILEPLPGM EFASHGDAYA YYQEYARSTG FNTAIQNSRR SKTSREFIDA 120 KFACSRYGTK REYEKFLNRP RSRQGGNLDP DNASGRRACS KTDCKASMHV KRRSDGKWII 180 HRFEKDHNHE LLPAQAVSEQ TRRMYAAMAR QFAEYKTAVC LNHDSRSQSE KSRNVAIDAE 240 AANSLIDFFV QMQSSFCNFF YAIDIGEDQR PRNFLWVDGK SRHDYGYFSD VISFDTSYVK 300 NKYKMPLALF VGVNQHYQFM LLGCALLSDE STSTFSWVMK NWLKAMGGQP PKIIITDQDK 360 GMKPAVSEVF PSTLHYFGLW QIFGKVSESL SYVIKQNESF MPKLEKCVYR SWTEEEFDRR 420 WNKLVERFGL KENELFRSLY EDRSRWVPNI MKDGFFAGMA SGQRSESVNS FFDKYVHRKT 480 TLQEFMKQYE AILQDRYEEE VKAASDTWNK QPAMKSPSPI EKHVAGIYTN AVFRKFQVEV 540 LGAVACMPKG EDQVGTAVKF RVHDFDMNQE FIVTLNEPES EICCICRLFE FRGFLCRHAM 600 LVLQIRGIST IPYRYILKRW TKDAKTGFSP GEGTENPQSR FQRFNDLCHK AMKLSEEGSL 660 SQESYRLTVR ALDDAFENCS NSNKNLHEAA SPGVLCIEED LQSGSLNKSN KKKASIKKRK 720 VNMEPEVMPV CSHETLQQMD KMSARTVSID GFFGHQPTSV QGMVRKN |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
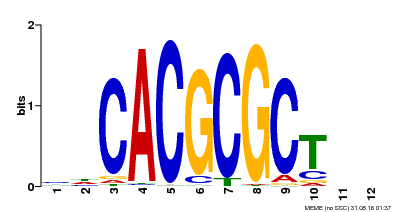
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011100397.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X2 | ||||
| Swissprot | Q9LIE5 | 0.0 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | S8BSB6 | 0.0 | S8BSB6_9LAMI; Uncharacterized protein (Fragment) | ||||
| STRING | Migut.M00094.1.p | 0.0 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA496 | 21 | 103 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 0.0 | far-red elongated hypocotyls 3 | ||||



