 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cotton_A_37559_BGI-A2_v1.0 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 836aa MW: 95830.2 Da PI: 7.2531 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 85.9 | 6.2e-27 | 84 | 186 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkk............tekerrtraetrtgCkaklkvkke 71
+fY+eYA+++GF++ +++s++sk+++e+++++f Cs++g+++e +k+ e++ +r+ ++t+Cka+++vk++
Cotton_A_37559_BGI-A2_v1.0 84 SFYQEYARSMGFNTAIQNSRRSKTSREFIDAKFACSRYGTKREYDKSfnrprarqskqdPENATGRRSCSKTDCKASMHVKRR 166
5*******************************************9999999988876655555559999************** PP
FAR1 72 kdgkwevtkleleHnHelap 91
dgkw+++++++eHnHel p
Cotton_A_37559_BGI-A2_v1.0 167 PDGKWVIHSFVKEHNHELLP 186
*****************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 9.6E-25 | 84 | 186 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 8.2E-29 | 285 | 376 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.507 | 564 | 600 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 1.8E-7 | 575 | 602 | IPR006564 | Zinc finger, PMZ-type |
| Pfam | PF04434 | 8.2E-4 | 575 | 597 | IPR007527 | Zinc finger, SWIM-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 836 aa Download sequence Send to blast |
MDIDLRLPSG EQCKEDEDAN GIDNMLDGDE SLHNGMVQVV GEDMRAEDGG EMHSSAVDMV 60 TFKEDTNLEP LSGMEFESHG EAYSFYQEYA RSMGFNTAIQ NSRRSKTSRE FIDAKFACSR 120 YGTKREYDKS FNRPRARQSK QDPENATGRR SCSKTDCKAS MHVKRRPDGK WVIHSFVKEH 180 NHELLPAQAV SEQTRRMYAA MARQFAEYKN VVGLKNDPKN PFDKGRNLAL EAADVKILLE 240 FFTHMQSINS NFFYAVDLGE DQRLKSLFWV DSKSRHDYSY FCDAVSFDTT YVTNKYKMPL 300 ALFIGVNHHY QCIPLGCALV SDESAATFSW LMKTWLKAMG GQSPRVLITD QDRTLKSVVA 360 EIFPNTLHCF FLWHVLGKVS ENLGHVIKQH GNLMAKFEKC IYRSWTDEEF GKRWWKILDR 420 FELKDDEWMK SLYEDRKKWV PTYIMNVLLA GMSTVQRAES VNSFFDKYVH KKTTVQDFLK 480 HYEAILQDRY EEEAKANSDS WSKVPALKSP SPFEKSVAGV YTHTLFKKFQ IEVVGAIACH 540 PKPENHDSTC SIFRVQDLEK NQDFIVTLNE MKAEVSCICR LYEYKGYLCR HAMVVLQING 600 HSAIPSQYIL KRWTKEAKSR HFMSEESEQV QSRVQRYNDL FQRGMKLIEE GSLSQESYYI 660 AFRSLEEAFG SCLSANTSNK SLAEAVASPT QGMICIEEDN QSRSTSKMNK KKNPTKKRKG 720 NSEQEVMAVA PTDGLQQMDK LSSRSVALDS YFGAQPSVQG MVQLNLMAPR DNYYGNQQTI 780 QGLLNTIATS HDGYYGTQQA MPGMGPMDFF RSPSFYIRDD PNVRVAQLHD DASRHT |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
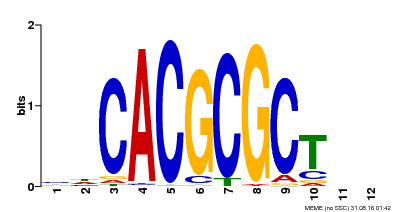
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017630678.1 | 0.0 | PREDICTED: protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| Swissprot | Q9LIE5 | 0.0 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A0D2PSJ9 | 0.0 | A0A0D2PSJ9_GOSRA; Uncharacterized protein | ||||
| STRING | Gorai.005G134200.1 | 0.0 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM118 | 26 | 253 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 0.0 | far-red elongated hypocotyls 3 | ||||



