 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cotton_A_29620_BGI-A2_v1.0 | ||||||||
| Common Name | F383_06989 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 406aa MW: 44148.7 Da PI: 9.9902 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 43.9 | 5.3e-14 | 328 | 379 | 5 | 56 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkev 56
+r+rr++kNRe+A rsR+RK+a++ eLe +v++L++ N++L + ee ++
Cotton_A_29620_BGI-A2_v1.0 328 RRQRRMIKNRESAARSRARKQAYTLELEAEVAKLKEMNEELLRKQEEVMEMQ 379
79************************************99988888877765 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 1.5E-12 | 324 | 390 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 10.645 | 326 | 377 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 5.13E-10 | 328 | 377 | No hit | No description |
| Gene3D | G3DSA:1.20.5.170 | 3.2E-13 | 328 | 378 | No hit | No description |
| Pfam | PF00170 | 3.0E-12 | 328 | 380 | IPR004827 | Basic-leucine zipper domain |
| CDD | cd14707 | 2.00E-25 | 328 | 382 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 331 | 346 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 406 aa Download sequence Send to blast |
MGSNLNFKRF GEAPSMEGSG SKAVGNFPLA RQSSIYSLTF DELQNTFGGL GKDFGSMNMD 60 ELLRNISTAE ETQGLMTASV PGGEGVSGGN LQRQGSLTLP RTLSQKTVEE VWKDLFKEND 120 GAKNVSNGGG GGGANLPQRQ QTLREMTLEE FLGRAGVVRE DMQPVGVPNN NGFFDNNSGL 180 ALQFQQTNGN NGFLSNNNSV LNQPPILPLD VSGAKSSHSQ QKQQPLFPKQ QTVAFAPSMH 240 LINTTHFPSP GARGSVVETS DLSMNTNLVQ SSGLQSGGMG IVGLPSPASH ISPDVISKNS 300 VDTTLSPVPY VLGRGRKRSA ALEKVVERRQ RRMIKNRESA ARSRARKQAY TLELEAEVAK 360 LKEMNEELLR KQEEVMEMQK NQMLETLNPA WGGKRQCLRR TLTGPW |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds to the ABA-responsive element (ABRE). Mediates stress-responsive ABA signaling. {ECO:0000269|PubMed:11884679, ECO:0000269|PubMed:15361142}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
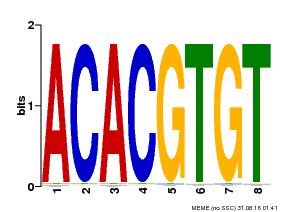
| Motif ID | Method | Source | Motif file |
| MP00038 | PBM | Transfer from AT4G34000 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by drought, salt, abscisic acid (ABA). {ECO:0000269|PubMed:10636868, ECO:0000269|PubMed:16284313}. | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | JX964384 | 1e-104 | JX964384.1 Gossypium hirsutum clone NBRI_TRANS-653 microsatellite sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017644690.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 7 | ||||
| Refseq | XP_017644691.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 7 | ||||
| Refseq | XP_017644692.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 7 | ||||
| Refseq | XP_017644693.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 7 | ||||
| Swissprot | Q9M7Q3 | 1e-119 | AI5L6_ARATH; ABSCISIC ACID-INSENSITIVE 5-like protein 6 | ||||
| TrEMBL | A0A0B0NJ24 | 0.0 | A0A0B0NJ24_GOSAR; Abscisic acid-insensitive 5-like protein 4 | ||||
| STRING | Gorai.013G044500.1 | 0.0 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM734 | 27 | 129 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G49720.1 | 3e-88 | abscisic acid responsive element-binding factor 1 | ||||



