 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cotton_A_27084_BGI-A2_v1.0 | ||||||||
| Common Name | F383_08377 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 275aa MW: 31300.2 Da PI: 6.6726 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 56.2 | 5.9e-18 | 79 | 131 | 4 | 56 |
-SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 4 RttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+ +++ eq+++Le+ Fe +++ e++ +LAk lgL+ rq+ +WFqNrRa++k
Cotton_A_27084_BGI-A2_v1.0 79 KKRLNLEQVKALEKSFELGNKLEPERKVQLAKALGLQPRQIAIWFQNRRARWK 131
4568889*********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 128.2 | 3.4e-41 | 77 | 167 | 1 | 91 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLeke 83
ekk+rl+ eqvk+LE+sFe +kLeperKv+la++Lglqprq+a+WFqnrRAR+ktkqlEkdy+aLk++++alk++n++L+++
Cotton_A_27084_BGI-A2_v1.0 77 EKKKRLNLEQVKALEKSFELGNKLEPERKVQLAKALGLQPRQIAIWFQNRRARWKTKQLEKDYDALKKQFEALKADNDALQAQ 159
69********************************************************************************* PP
HD-ZIP_I/II 84 veeLreel 91
+++L++el
Cotton_A_27084_BGI-A2_v1.0 160 NKKLNAEL 167
*****998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.46E-19 | 69 | 135 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 16.763 | 73 | 133 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.0E-17 | 76 | 137 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.65E-16 | 78 | 134 | No hit | No description |
| Pfam | PF00046 | 3.3E-15 | 79 | 131 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 5.1E-19 | 81 | 141 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 9.4E-6 | 104 | 113 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 108 | 131 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 9.4E-6 | 113 | 129 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 4.4E-14 | 133 | 173 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 275 aa Download sequence Send to blast |
MAFPPHAFMF QPHEDHHNDH LPSPTSLNFL PSCPPQLFHG GGAPFMMKRS VSFSGVDKSE 60 EVHGDDELSD DGSHLGEKKK RLNLEQVKAL EKSFELGNKL EPERKVQLAK ALGLQPRQIA 120 IWFQNRRARW KTKQLEKDYD ALKKQFEALK ADNDALQAQN KKLNAELLAL KTKDSNETSC 180 IKKENDCSWS YGSDNNSCDV NLDISRTPLM SSSKHLFPPS VRPTSMTQLL QGSSRPDLQC 240 VKLDQVVQEE SFCNMFNGVD EQQAFWPWSE QQSFH |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 125 | 133 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00322 | DAP | Transfer from AT3G01220 | Download |
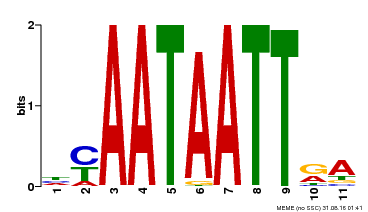
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. {ECO:0000269|PubMed:12644682}. | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | EF151309 | 0.0 | EF151309.1 Gossypium hirsutum homeobox protein (HB1) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017609830.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein HAT7-like | ||||
| Swissprot | Q8LAT0 | 3e-97 | ATB20_ARATH; Homeobox-leucine zipper protein ATHB-20 | ||||
| TrEMBL | A0A0B0PXN9 | 0.0 | A0A0B0PXN9_GOSAR; Homeobox-leucine zipper ATHB-20-like protein | ||||
| STRING | Gorai.006G022100.1 | 0.0 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3095 | 26 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G01220.1 | 7e-91 | homeobox protein 20 | ||||



