 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cotton_A_23397_BGI-A2_v1.0 | ||||||||
| Common Name | F383_07659 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 685aa MW: 75643 Da PI: 6.7857 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 74.1 | 1.6e-23 | 121 | 222 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--T CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLke 76
f k+lt+sd+++ g +++p+ +ae++ +++ + t+ +d +g++W++++iyr++++r++lt+GW++Fv+ ++L +
Cotton_A_23397_BGI-A2_v1.0 121 FAKTLTQSDANNGGGFSVPRYCAETIfprldynAEPPVQ--TILAKDVHGEVWKFRHIYRGTPRRHLLTTGWSNFVNHKKLVA 201
89****************************985444444..8999************************************** PP
T-EEEEEE-SSSEE..EEEEE-S CS
B3 77 gDfvvFkldgrsefelvvkvfrk 99
gD++vF + +++ l+v+++r+
Cotton_A_23397_BGI-A2_v1.0 202 GDSIVFLR--ADNGDLCVGIRRA 222
*******3..477888****996 PP
| |||||||
| 2 | Auxin_resp | 112.5 | 4.1e-37 | 281 | 364 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrl.sGtvvgvsdldpvrWpnSkWrsL 82
aa+ a+++++FevvY+Pr+st+eF+vk+++v++a+++++ +GmRfkm fetedss++++ +Gt++ + +dp+rWpnS+Wr+L
Cotton_A_23397_BGI-A2_v1.0 281 AASHAASGQPFEVVYYPRTSTPEFCVKASSVRAAMQIQWYPGMRFKMPFETEDSSRISWfMGTISTAQVVDPIRWPNSPWRLL 363
678899****************************************************************************9 PP
Auxin_resp 83 k 83
+
Cotton_A_23397_BGI-A2_v1.0 364 Q 364
7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 5.1E-29 | 113 | 224 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 1.1E-36 | 116 | 224 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 8.50E-22 | 120 | 221 | No hit | No description |
| Pfam | PF02362 | 1.5E-21 | 121 | 222 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 6.6E-22 | 121 | 223 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.183 | 121 | 223 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 2.0E-32 | 281 | 364 | IPR010525 | Auxin response factor |
| PROSITE profile | PS51745 | 20.044 | 599 | 682 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007389 | Biological Process | pattern specification process | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0048829 | Biological Process | root cap development | ||||
| GO:0051301 | Biological Process | cell division | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 685 aa Download sequence Send to blast |
MITVMDSRKE VVKNSEKCLD PQLWHACAGG MVQMPSVNSK VFYFPQGHAE HANGNVDFGN 60 LPIPSLVLCR VSAVRFMADP ETDEVYAKIM LIPLRENSFG VEDDGFDGNV GVENPEKSAS 120 FAKTLTQSDA NNGGGFSVPR YCAETIFPRL DYNAEPPVQT ILAKDVHGEV WKFRHIYRGT 180 PRRHLLTTGW SNFVNHKKLV AGDSIVFLRA DNGDLCVGIR RAKRGLGGGH EFPGWNSVSG 240 TSGSQVGSYS PFLREGESKL MRKDCNGDPR GRIRADSVIE AASHAASGQP FEVVYYPRTS 300 TPEFCVKASS VRAAMQIQWY PGMRFKMPFE TEDSSRISWF MGTISTAQVV DPIRWPNSPW 360 RLLQVAWDEP DLLHNVKRVS PWLVELVTNI PAINLNPFSP PRKKMRLPQH PDISFLNQIP 420 MPSFSGNSFR SSSPMRCITD NIPGGIQGAR HEPFGLSSSD LRSNKLHSGL FPSGFHQLDR 480 TAPPTRLSGN NFCSDNANNT NISSLLTIGN PTQSFKQSND SKTPHIVLFG QLIFCEQRAS 540 QSCSGDTVGN SSSDGNTEKT AISSDGTGSV LHQNVRENSS DEGFPWCKED QKTDLGLETG 600 HCKVFMESEN VGRTLDLSVL GSYEELYGKL ANMFGIESSE MLSSVLYRDA AGSVKHTGDE 660 PFSEFMKTAR RLTILMDSSS DNLER |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldu_A | 4e-78 | 1 | 385 | 33 | 388 | Auxin response factor 5 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
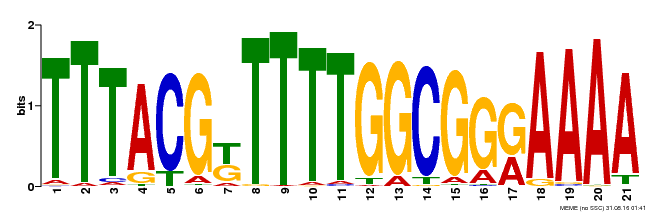
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00461 | DAP | Transfer from AT4G30080 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017649647.1 | 0.0 | PREDICTED: auxin response factor 18-like isoform X2 | ||||
| Swissprot | Q653H7 | 0.0 | ARFR_ORYSJ; Auxin response factor 18 | ||||
| TrEMBL | A0A0B0PKQ8 | 0.0 | A0A0B0PKQ8_GOSAR; Auxin response factor | ||||
| STRING | Gorai.011G238900.1 | 0.0 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM580 | 27 | 130 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G30080.1 | 0.0 | auxin response factor 16 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



