 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cotton_A_11128_BGI-A2_v1.0 | ||||||||
| Common Name | F383_29826 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 471aa MW: 50893.5 Da PI: 7.0814 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 101.3 | 6.2e-32 | 250 | 305 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+p+LH+rFv+a++ LGGs++AtPk+i+elmkv+gLt+++vkSHLQkYRl+
Cotton_A_11128_BGI-A2_v1.0 250 KARRCWSPDLHRRFVNALQMLGGSQVATPKQIRELMKVEGLTNDEVKSHLQKYRLH 305
68****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 11.407 | 247 | 307 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.15E-16 | 248 | 308 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 3.9E-28 | 248 | 308 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.5E-26 | 250 | 305 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 8.4E-7 | 252 | 303 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009266 | Biological Process | response to temperature stimulus | ||||
| GO:0009740 | Biological Process | gibberellic acid mediated signaling pathway | ||||
| GO:0010452 | Biological Process | histone H3-K36 methylation | ||||
| GO:0048579 | Biological Process | negative regulation of long-day photoperiodism, flowering | ||||
| GO:1903507 | Biological Process | negative regulation of nucleic acid-templated transcription | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 471 aa Download sequence Send to blast |
MASPTGLTLD CKPHSYSMLL KSFGDQQIDQ TQKLEDFLGR LEEERLKIDA FKRELPLCMQ 60 LLTNAVEASR QQLHACRANQ GSRPVLEEFI PLKNSSSENS DKSQNISDKA NWMTSAQLWS 120 QAGNETKPPQ SSVASPKEAD IGFNVSPKLG LDTKQRNGGA FLPFSKDRNN PCPGSTLQPL 180 PDLALASMNE DMDDKKCSEA ENGCQRRENS GKTGNGGALV EQGKGTSCNA AEGQTTNGNT 240 NTNTGQPHRK ARRCWSPDLH RRFVNALQML GGSQVATPKQ IRELMKVEGL TNDEVKSHLQ 300 KYRLHTRRPS PSPQAPGAPA PQLVVLGGIW VSPEYATAAA AAHGGAPTLY GAHHPVASHA 360 PPPHFCASPV PQEFYSTTAT PASTPPPQLH HHTLQQQLHM YKSTSQAHSL PESDVRGAGD 420 RSESIEDGKS ESSSWKGESG ENGGGADQRK GLAALREDGE ESNGSEITLK F |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 1e-14 | 250 | 303 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 1e-14 | 250 | 303 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 1e-14 | 250 | 303 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 1e-14 | 250 | 303 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor acting as a flowering repressor, directly repressing FT expression in a dosage-dependent manner in the leaf vasculature. {ECO:0000269|PubMed:25132385}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00261 | DAP | Transfer from AT2G03500 | Download |
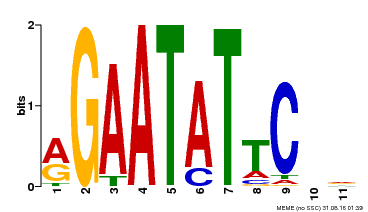
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by SVP. Down-regulated by high temperature and gibberellic acid treatment. Not regulated by photoperiod, circadian rhythm under long days or vernalization. {ECO:0000269|PubMed:25132385}. | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | JX615064 | 2e-32 | JX615064.1 Gossypium hirsutum clone NBRI_GE58863 microsatellite sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017610040.1 | 0.0 | PREDICTED: myb family transcription factor EFM | ||||
| Swissprot | Q9ZQ85 | 1e-137 | EFM_ARATH; Myb family transcription factor EFM | ||||
| TrEMBL | A0A0B0MUB9 | 0.0 | A0A0B0MUB9_GOSAR; Two-component response regulator ARR18-like protein | ||||
| STRING | Gorai.009G022100.1 | 0.0 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM7134 | 28 | 43 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G03500.1 | 1e-110 | G2-like family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



