 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cotton_A_05396_BGI-A2_v1.0 | ||||||||
| Common Name | F383_08946 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 838aa MW: 92035.5 Da PI: 6.3815 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 58.4 | 1.2e-18 | 17 | 75 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++++++ps +r++L +++ +++ +q+kvWFqNrR +ek+
Cotton_A_05396_BGI-A2_v1.0 17 KYVRYTPEQVEALERLYHECPKPSSIRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 75
5679*****************************************************97 PP
| |||||||
| 2 | START | 181 | 6.9e-57 | 161 | 369 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEE CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlev 84
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s++++g a+ra+g+v +++ v+ell+d++ W ++++++++l+v
Cotton_A_05396_BGI-A2_v1.0 161 IAEETLAEFLSKATGTAVEWVQMPGMKPGPDSIGIIAISHGCPGVAARACGLVGLEPT-RVAELLKDRPSWFHDCRAVDVLNV 242
7899******************************************************.9*********************** PP
ECTT..EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCE CS
START 85 issg..galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksngh 161
+ ++ g+++l +++l+a+++l+p Rdf+ +Ry+ l++g++v++++S+ ++q+ p+ +++vRae+lpSg+li+p+++g+
Cotton_A_05396_BGI-A2_v1.0 243 LPTAngGTIELLYMQLYAPTTLAPaRDFWLLRYTSVLEDGSLVVCERSLKNTQNGPSmppVQHFVRAEMLPSGYLIRPCEGGG 325
**9999*************************************************9988899********************* PP
EEEEEEE-EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXX CS
START 162 skvtwvehvdlkgrlphwllrslvksglaegaktwvatlqrqce 205
s +++v+h+dl+ ++++++lr+l++s+++ ++kt++a+l+++++
Cotton_A_05396_BGI-A2_v1.0 326 SIIHIVDHMDLEPWSVPEVLRPLYESSTVLAQKTTMAALRQLRQ 369
****************************************9876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.613 | 12 | 76 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.2E-15 | 14 | 80 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 3.38E-17 | 16 | 80 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 9.31E-17 | 17 | 77 | No hit | No description |
| Pfam | PF00046 | 2.9E-16 | 18 | 75 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 1.6E-18 | 19 | 75 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 1.34E-6 | 69 | 108 | No hit | No description |
| PROSITE profile | PS50848 | 25.339 | 151 | 366 | IPR002913 | START domain |
| CDD | cd08875 | 9.97E-78 | 155 | 371 | No hit | No description |
| SMART | SM00234 | 6.5E-43 | 160 | 370 | IPR002913 | START domain |
| Gene3D | G3DSA:3.30.530.20 | 2.1E-23 | 160 | 367 | IPR023393 | START-like domain |
| SuperFamily | SSF55961 | 4.26E-39 | 160 | 372 | No hit | No description |
| Pfam | PF01852 | 2.8E-54 | 161 | 369 | IPR002913 | START domain |
| Pfam | PF08670 | 3.7E-53 | 695 | 837 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 838 aa Download sequence Send to blast |
MAMSCKDGKP GSLDNGKYVR YTPEQVEALE RLYHECPKPS SIRRQQLIRE CPILSNIEPK 60 QIKVWFQNRR CREKQRKEAS RLQAVNRKLT AMNKLLMEEN DRLQKQVSQL VYENGCFRQH 120 TQNATLATKD PSCESAVMSG QQRETHQHPP RDASPAGLLS IAEETLAEFL SKATGTAVEW 180 VQMPGMKPGP DSIGIIAISH GCPGVAARAC GLVGLEPTRV AELLKDRPSW FHDCRAVDVL 240 NVLPTANGGT IELLYMQLYA PTTLAPARDF WLLRYTSVLE DGSLVVCERS LKNTQNGPSM 300 PPVQHFVRAE MLPSGYLIRP CEGGGSIIHI VDHMDLEPWS VPEVLRPLYE SSTVLAQKTT 360 MAALRQLRQI AQEVSQSNVT GWGRRPAALR ALSQRLSRGF NDAVNGFTDE GWSMMGNDGM 420 DDVTILVNSS PDKLMGLNLS FANGFPSVSN AVLCAKASML LQNVPPAILL RFLREHRSEW 480 ADSSIDAYSA AVVKVGPCSL RGSRVGGFGS QVILPLAHTV EHEEFLEVIK LEGIAQSPED 540 ALMPRDVFLL QLCSGMDENA VGTCAELIFA PIDASFADDA PLLPSGFRII PLDSGKEASS 600 PNRTLDLASA LEIGPAGNKA SNDYSAAGGC MRSVMTIAFE FAFESHLQEH VASMARQYVR 660 SIISSVQRVA LALSPSYSGS HAGLRTPLGT PEAQTLARWI CQSYRVYMGV ELLKSGTEGG 720 ESVLKTLWHH SDAIMCCSLK ALPVFTFANQ AGLDMLETTL VALQDLTLEK IFDEHGRKTL 780 CTEFPQIMQQ GFACLQGGIC LSSMGRPVSY ERAVAWKVLN EEENAHCICF MFVNWSFV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
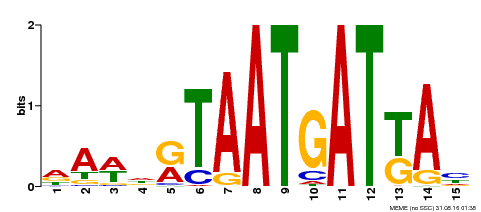
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012443274.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-15 | ||||
| Refseq | XP_017606589.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-15 | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | A0A0B0NUV3 | 0.0 | A0A0B0NUV3_GOSAR; Homeobox-leucine zipper ATHB-15-like protein | ||||
| TrEMBL | A0A0D2QFF5 | 0.0 | A0A0D2QFF5_GOSRA; Uncharacterized protein | ||||
| STRING | Gorai.009G136800.1 | 0.0 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2022 | 26 | 78 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



