 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | mrna19228.1-v1.0-hybrid | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Rosoideae; Potentilleae; Fragariinae; Fragaria
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 386aa MW: 43219.8 Da PI: 7.1502 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 41 | 4.5e-13 | 70 | 114 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT +E++++++a +++ + Wk+I + +g +t q++s+ qky
mrna19228.1-v1.0-hybrid 70 RESWTDQEHDKFLEALQLFDRD-WKKIEAFVG-SKTVIQIRSHAQKY 114
789*****************77.*********.*************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.5E-16 | 64 | 120 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 15.014 | 65 | 119 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.8E-6 | 68 | 120 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 3.2E-18 | 68 | 117 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 1.4E-10 | 69 | 117 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.2E-10 | 70 | 114 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 7.49E-8 | 72 | 115 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 386 aa Download sequence Send to blast |
MVSVNPNPAP QGFYLFDPTN MGLPGLNSMP PPTTPSTTST TMSSAAQNAA SLAEDQSKKI 60 RKPYTITKSR ESWTDQEHDK FLEALQLFDR DWKKIEAFVG SKTVIQIRSH AQKYFLKVQK 120 KGTSEHVPPP RPKRKATHPY PQKAPKSAPV VSQVVGPFES SSALLEHGYT YQPDSSFVHS 180 TPVTSATLSS WSHSYVPPVN ASQKTKDDGR LAGSADAQNS CYSSSNESKP TTWSMGETFG 240 RACHHQPQRV LPDFAQVYKF IGSVFDPDST CHLERLKQMD PINLETALLL MRNLSINLSS 300 PEFENHYGKD YEFEAPVELL DKMLHSLRQS PDEQLVVVSQ VLVFDINIWY EDIVNTQVLA 360 FNGKPVKNLK SLARLVENCE DEFLKI |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5il9_A | 8e-27 | 298 | 385 | 436 | 523 | Protease Do-like 9 |
| 5il9_B | 8e-27 | 298 | 385 | 436 | 523 | Protease Do-like 9 |
| 5ilb_A | 7e-27 | 298 | 385 | 344 | 431 | Protease Do-like 2, chloroplastic,Protease Do-like 9 |
| 5ilb_B | 7e-27 | 298 | 385 | 344 | 431 | Protease Do-like 2, chloroplastic,Protease Do-like 9 |
| 5jyk_A | 8e-27 | 298 | 385 | 436 | 523 | Protease Do-like 9 |
| 5jyk_B | 8e-27 | 298 | 385 | 436 | 523 | Protease Do-like 9 |
| 5y09_A | 8e-27 | 298 | 385 | 436 | 523 | Protease Do-like 9 |
| 5y09_B | 8e-27 | 298 | 385 | 436 | 523 | Protease Do-like 9 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
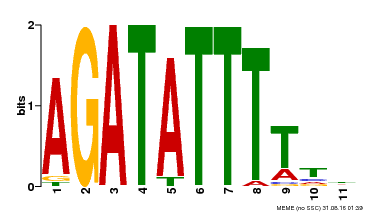
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00423 | DAP | Transfer from AT4G01280 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | mrna19228.1-v1.0-hybrid |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004306739.1 | 0.0 | PREDICTED: protein REVEILLE 3-like | ||||
| Swissprot | C0SVG5 | 1e-100 | RVE5_ARATH; Protein REVEILLE 5 | ||||
| TrEMBL | A0A2P6SHV1 | 0.0 | A0A2P6SHV1_ROSCH; Putative transcription factor MYB-HB-like family | ||||
| STRING | XP_004306739.1 | 0.0 | (Fragaria vesca) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1119 | 33 | 96 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G01280.2 | 1e-101 | MYB_related family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | mrna19228.1-v1.0-hybrid |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



