 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | mrna05518.1-v1.0-hybrid | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Rosoideae; Potentilleae; Fragariinae; Fragaria
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 489aa MW: 53312.3 Da PI: 5.8751 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 105.2 | 3.7e-33 | 255 | 309 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
+pr+rWtpeLHerFveav++L G+ekAtPk++l++m+v+gLt++hvkSHLQkYRl
mrna05518.1-v1.0-hybrid 255 RPRMRWTPELHERFVEAVKKLDGAEKATPKAVLKVMNVEGLTIYHVKSHLQKYRL 309
59****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 11.007 | 252 | 312 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.56E-17 | 254 | 309 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 4.1E-31 | 254 | 310 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 5.4E-25 | 256 | 310 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 4.8E-10 | 257 | 308 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 2.7E-23 | 345 | 392 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0007134 | developmental stage | sporophyte vegetative stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 489 aa Download sequence Send to blast |
MKVGLSSAIM SLHGVSSVTQ SGTTKGITQS YCTSLSPAHD FLGCESEGRN SVAHECSSTR 60 LSPFMRTESF SSPTNMRESS LQRVKSTFSR SSVFCTSLYQ SSSSTSETSR QLGNLPFLPH 120 PPTYSQSNSA VDSTSPLLLS QDMSNQYDDE QSDYLMKDFL NMTGDASDGS FHEIGCGSDT 180 MALTEQLEFQ FLSDQLDIAI TDNGENPRLD EIYDIPRASS EPTIELTCSK SCGSTAPLVD 240 ALSSHPSPGP SSAHRPRMRW TPELHERFVE AVKKLDGAEK ATPKAVLKVM NVEGLTIYHV 300 KSHLQKYRLA KYMPEKKEDK KASSSEEKKA ASSGNESDGR RKGSIHITEA LRMQMEVQKQ 360 LHEQLEVQRS LQLRIEEHAK YLEKILEEQQ KAGSALLSPQ ALSSLTTNSL KDSEQQPPPS 420 ACISASQPAA SDSSSPDSPL SLKHKAAACS ESEAHAYTKK LRIEEKPDDP VVENPSATVT 480 TTPTTTSQ* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 9e-25 | 255 | 312 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 9e-25 | 255 | 312 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 9e-25 | 255 | 312 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 9e-25 | 255 | 312 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 9e-25 | 255 | 312 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 9e-25 | 255 | 312 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 9e-25 | 255 | 312 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 9e-25 | 255 | 312 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00354 | DAP | Transfer from AT3G13040 | Download |
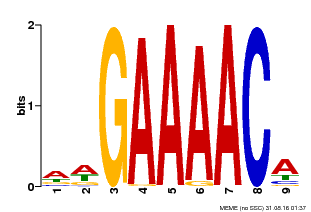
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | mrna05518.1-v1.0-hybrid |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011467504.1 | 0.0 | PREDICTED: protein PHR1-LIKE 1 | ||||
| Refseq | XP_011467505.1 | 0.0 | PREDICTED: protein PHR1-LIKE 1 | ||||
| Swissprot | Q949U2 | 1e-148 | PHL6_ARATH; Myb family transcription factor PHL6 | ||||
| TrEMBL | A0A2P6RZF1 | 0.0 | A0A2P6RZF1_ROSCH; Putative transcription factor MYB-HB-like family | ||||
| STRING | XP_004303787.1 | 0.0 | (Fragaria vesca) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF13469 | 19 | 25 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G13040.2 | 1e-117 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | mrna05518.1-v1.0-hybrid |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



