 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | mrna02647.1-v1.0-hybrid | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Rosoideae; Potentilleae; Fragariinae; Fragaria
|
||||||||
| Family | TALE | ||||||||
| Protein Properties | Length: 478aa MW: 54344.9 Da PI: 8.0553 | ||||||||
| Description | TALE family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 28.2 | 3.2e-09 | 347 | 381 | 21 | 55 |
HSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHH CS
Homeobox 21 knrypsaeereeLAkklgLterqVkvWFqNrRake 55
k +yps++e+ LA+++gL+++q+ +WF N+R ++
mrna02647.1-v1.0-hybrid 347 KWPYPSESEKVALAESTGLDQKQINNWFINQRKRH 381
669*****************************885 PP
| |||||||
| 2 | ELK | 37.5 | 5.2e-13 | 301 | 322 | 1 | 22 |
ELK 1 ELKhqLlrKYsgyLgsLkqEFs 22
ELK++LlrKYsgyL+sLkqE+s
mrna02647.1-v1.0-hybrid 301 ELKNHLLRKYSGYLSSLKQELS 322
9*******************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01255 | 3.3E-20 | 114 | 158 | IPR005540 | KNOX1 |
| Pfam | PF03790 | 1.2E-22 | 115 | 156 | IPR005540 | KNOX1 |
| SMART | SM01256 | 3.5E-26 | 166 | 217 | IPR005541 | KNOX2 |
| Pfam | PF03791 | 1.5E-23 | 171 | 215 | IPR005541 | KNOX2 |
| SMART | SM01188 | 3.5E-7 | 301 | 322 | IPR005539 | ELK domain |
| Pfam | PF03789 | 3.5E-10 | 301 | 322 | IPR005539 | ELK domain |
| PROSITE profile | PS51213 | 11.296 | 301 | 321 | IPR005539 | ELK domain |
| PROSITE profile | PS50071 | 12.957 | 321 | 384 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 2.31E-19 | 322 | 396 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 2.8E-13 | 323 | 388 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 2.5E-27 | 326 | 387 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 2.72E-11 | 333 | 385 | No hit | No description |
| Pfam | PF05920 | 1.5E-16 | 341 | 380 | IPR008422 | Homeobox KN domain |
| PROSITE pattern | PS00027 | 0 | 359 | 382 | IPR017970 | Homeobox, conserved site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001708 | Biological Process | cell fate specification | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010051 | Biological Process | xylem and phloem pattern formation | ||||
| GO:0010089 | Biological Process | xylem development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 478 aa Download sequence Send to blast |
MEEYNNSDEM DHESQGTPRG NFLYASPTLG GNYGRAAAAS DHNQQNNTFH LQSSGGECFP 60 STQSTVKTEA AGSTSQHARH KFHHPQFPPP SLVLSRGHND QQLPVQQQNE NEVEAIKAKI 120 IAHPHYSNLL QAYMDCQRVG APSEVVARLS AARQEFVARQ RSSVSSRDAS SKDPELDQFM 180 EAYYDMLVKY REELTRPIQE AMDFMKKIET QLNMLGNNST ATPLRIFSAS ALAGVVPAGL 240 SHYKIHPLLV FTRRQKVRNL DEKHSTYDKC DGNGSSEEDQ DNNSGGETEV AEIDPRAEDR 300 ELKNHLLRKY SGYLSSLKQE LSKKKKKGKL PKDARQKLLS WWELHYKWPY PSESEKVALA 360 ESTGLDQKQI NNWFINQRKR HWKPSEDMQF MVMDGLHPQN AAALYMDGHY IGDGHYRLEK 420 VKRLVKWTFW DLTLTLTVKA SMREKCYTIR DYDFNAAFQV GMRLGYLDQP NGGLGPK* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably binds to the DNA sequence 5'-TGAC-3'. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00675 | ChIP-seq | Transfer from GRMZM2G017087 | Download |
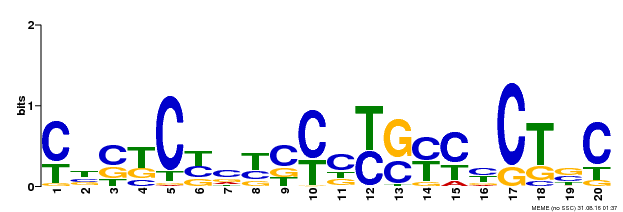
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | mrna02647.1-v1.0-hybrid |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HQ413775 | 0.0 | HQ413775.1 Fragaria vesca knotted-like homeobox KNOX2 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001266972.1 | 0.0 | homeobox protein knotted-1-like 2-like | ||||
| Swissprot | O04135 | 0.0 | KNAP2_MALDO; Homeobox protein knotted-1-like 2 | ||||
| TrEMBL | F5A6B2 | 0.0 | F5A6B2_FRAVE; Knotted-like homeobox KNOX2 | ||||
| STRING | XP_004291007.1 | 0.0 | (Fragaria vesca) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4584 | 32 | 46 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G08150.1 | 1e-128 | KNOTTED-like from Arabidopsis thaliana | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | mrna02647.1-v1.0-hybrid |



