 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | mrna02324.1-v1.0-hybrid | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Rosoideae; Potentilleae; Fragariinae; Fragaria
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 555aa MW: 60669 Da PI: 6.3824 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 49.8 | 8e-16 | 101 | 148 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+grWT+eEd+ l+++++ G g+W++ ++ g+ R++k+c++rw +yl
mrna02324.1-v1.0-hybrid 101 KGRWTAEEDQTLINYIRANGEGSWRSLPKNAGLLRCGKSCRLRWINYL 148
79******************************99************97 PP
| |||||||
| 2 | Myb_DNA-binding | 51.2 | 2.8e-16 | 154 | 199 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg+ T++E+++++++++ lG++ W++Ia+ ++ gRt++++k++w+++l
mrna02324.1-v1.0-hybrid 154 RGNITPQEEDIIIKLHASLGNR-WSLIASQLP-GRTDNEIKNYWNSHL 199
7999******************.*********.************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 2.1E-21 | 92 | 151 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 16.583 | 96 | 148 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 9.56E-28 | 98 | 195 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 8.7E-12 | 100 | 150 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.7E-14 | 101 | 148 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 9.27E-9 | 103 | 148 | No hit | No description |
| PROSITE profile | PS51294 | 25.801 | 149 | 203 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.0E-25 | 152 | 203 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 3.4E-16 | 153 | 201 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.0E-14 | 154 | 199 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.21E-10 | 158 | 199 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 555 aa Download sequence Send to blast |
MKEKSICPIN KGAGLAKAHS LRYIWQLRAC TAEGGIRSEV ETERGGRARR RDDWMSRVRA 60 LRTYMDVGNA KPHHEPLDNG EATTLTEMGR APCCEKVGLK KGRWTAEEDQ TLINYIRANG 120 EGSWRSLPKN AGLLRCGKSC RLRWINYLRA DLKRGNITPQ EEDIIIKLHA SLGNRWSLIA 180 SQLPGRTDNE IKNYWNSHLS RKIDAFRRPI NKIHMPSASA AAAASSSSST TTAADEINMA 240 MKLGPPSSKR RGRGGRTSRW AMKKNKVTYA STAINHHLDS CTHNHDDKSA TMTSSSSPPP 300 PSAAAAAVAA GTGQKETQAL VGGGPSDGSA AGKFDDGGEV AFNDLMMDVA NEIISDPTGV 360 LTLIGEKQND DGDDDTGVIR DDEDTGKLLC GPNKVMTNAT AAASHEEEHH QLDQISATNL 420 SSSSNSNSKS SYGEAEAAAF DGDDWYNSCS NYSPGSAAAA TCKFDQQKQQ RDGINGTVND 480 EMTWDWESDI HRYDYLWNVD QKDDENMLSW LWEQHAADDH HLDHSNKSRS KAVVVDDDCD 540 PADKHNAILA WLLS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 5e-24 | 99 | 204 | 5 | 109 | B-MYB |
| 1h8a_C | 1e-23 | 99 | 203 | 25 | 128 | MYB TRANSFORMING PROTEIN |
| Search in ModeBase | ||||||
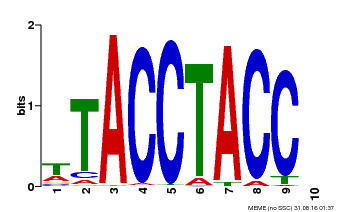
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00627 | PBM | Transfer from PK06182.1 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | mrna02324.1-v1.0-hybrid |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | DQ074470 | 1e-113 | DQ074470.1 Malus x domestica MYB22 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004301809.1 | 0.0 | PREDICTED: transcription factor MYB12-like | ||||
| TrEMBL | A0A2P6PIB4 | 0.0 | A0A2P6PIB4_ROSCH; Putative transcription factor MYB-HB-like family | ||||
| STRING | XP_004301809.1 | 0.0 | (Fragaria vesca) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF31 | 34 | 817 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G62610.1 | 3e-69 | myb domain protein 11 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | mrna02324.1-v1.0-hybrid |



