 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | mrna00431.1-v1.0-hybrid | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Rosoideae; Potentilleae; Fragariinae; Fragaria
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 696aa MW: 75754.7 Da PI: 5.0833 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 56.4 | 7e-18 | 172 | 219 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WTt Ed +lv++vk++G g+W+++ ++ g+ R++k+c++rw ++l
mrna00431.1-v1.0-hybrid 172 KGPWTTAEDAILVEYVKKHGEGNWNAVQKHSGLFRCGKSCRLRWANHL 219
79******************************************9986 PP
| |||||||
| 2 | Myb_DNA-binding | 53 | 8e-17 | 225 | 268 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g++T+eE+ l+v++++++G++ W++ a++++ gRt++++k++w++
mrna00431.1-v1.0-hybrid 225 KGAFTPEEERLIVELHAKMGNK-WARMAAHLP-GRTDNEIKNYWNT 268
799*******************.*********.***********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 16.678 | 167 | 219 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.3E-15 | 171 | 221 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 2.43E-30 | 171 | 266 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 1.1E-15 | 172 | 219 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 8.7E-24 | 173 | 226 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 7.06E-12 | 174 | 219 | No hit | No description |
| PROSITE profile | PS51294 | 26.711 | 220 | 274 | IPR017930 | Myb domain |
| SMART | SM00717 | 3.4E-17 | 224 | 272 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.1E-15 | 225 | 268 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.97E-12 | 227 | 268 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 6.2E-26 | 227 | 273 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0007134 | developmental stage | sporophyte vegetative stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 696 aa Download sequence Send to blast |
MLSLENITSD TKFHLITNRL RPFDETRNSP PHPDRHGTLS APTRTNAAAL RCVAPSTLAS 60 SPSSSTNSGV PLIRFSQLSD QFDTTLETIL ICDSRFPVLE IIDCEKGPIF WVFDSVKGQC 120 RVSACITSGS KEEMSRTTSD SEDGMFSKDQ IESPLMDENN GGIGNGGIIL KKGPWTTAED 180 AILVEYVKKH GEGNWNAVQK HSGLFRCGKS CRLRWANHLR PNLKKGAFTP EEERLIVELH 240 AKMGNKWARM AAHLPGRTDN EIKNYWNTRI KRRQRAGLPL YPPELSLQAS QESQQGHSGG 300 ALNGTDRAHH DSLQSNSYEI PDVVFDSLKG NNCVLPYVPE LPDISASGML MKGLGSSPYC 360 GFMPPTMHRQ KRPRESTALF PTSGSSFRDG FPQFDQYEND ACDKVARPFG LPFLPDPDPT 420 TKSPLSFGVI QGCHSLSNGN SSACRPTYGP VKLELPSLQY PETDLGSWST SPPPPLLESF 480 DNFIQSPPPV GAFESDCASP RNSGLLDALL YESKTLSSTK NHSTDKSSIS SSVTPGEVAD 540 SSTLNVCETE WEEYGDPISP LGHSATLLFS ECTPISASGS SLEEAPPAET ITGSNVKPEP 600 ADHAWTPEKQ KESSPPLDYI RPDALLGSDW LEPSTGIKDQ SIMNMNDAIV SLLGEDLVTE 660 YKHMASGTST SNQGWGLGSC PWNNMPAVCQ MTDLP* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 2e-32 | 170 | 273 | 5 | 107 | B-MYB |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
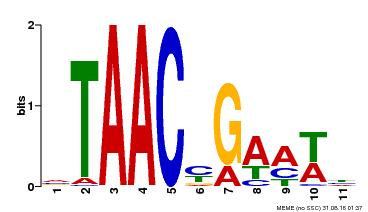
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | mrna00431.1-v1.0-hybrid |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011468770.1 | 0.0 | PREDICTED: transcription factor GAMYB | ||||
| TrEMBL | A0A2P6SB49 | 0.0 | A0A2P6SB49_ROSCH; Putative transcription factor MYB-HB-like family | ||||
| STRING | XP_004306467.1 | 0.0 | (Fragaria vesca) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2047 | 34 | 85 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.3 | 1e-133 | myb domain protein 33 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | mrna00431.1-v1.0-hybrid |



