 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | FANhyb_rscf00002218.1.g00001.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Rosoideae; Potentilleae; Fragariinae; Fragaria
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 735aa MW: 81181.9 Da PI: 7.3402 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 74.2 | 1.5e-23 | 121 | 222 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHH CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkan 72
f k+lt+sd+++ g +++p+ +ae++ ++ + t+ +d +g++W++++iyr++++r++lt+GW+ Fv+ +
FANhyb_rscf00002218.1.g00001.1 121 FAKTLTQSDANNGGGFSVPRYCAETIfprldysADPPVQ--TILAKDVHGETWKFRHIYRGTPRRHLLTTGWSTFVNHK 197
89****************************996555555..89************************************ PP
T--TT-EEEEEE-SSSEE..EEEEE-S CS
B3 73 gLkegDfvvFkldgrsefelvvkvfrk 99
+L +gD++vF + +++ l+v+++r+
FANhyb_rscf00002218.1.g00001.1 198 KLVAGDSIVFL--RAENGDLCVGIRRA 222
***********..4488999****997 PP
| |||||||
| 2 | Auxin_resp | 116.9 | 1.8e-38 | 288 | 371 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrl.sGtvvgvsdldpvrWpnSk 78
aa+ as++++FevvY+Prast+eF+vk++ v++al++++++GmRfkmafetedss++++ +Gt+++v+ +dp+rWp+S+
FANhyb_rscf00002218.1.g00001.1 288 AATLASNGQPFEVVYYPRASTPEFCVKASLVKAALQIRWCPGMRFKMAFETEDSSRISWfMGTISSVQVSDPMRWPESP 366
6899*************************************************************************** PP
Auxin_resp 79 WrsLk 83
Wr+L+
FANhyb_rscf00002218.1.g00001.1 367 WRLLQ 371
***97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 9.9E-38 | 113 | 225 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 2.62E-29 | 114 | 224 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 3.10E-21 | 120 | 221 | No hit | No description |
| Pfam | PF02362 | 9.5E-22 | 121 | 222 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.112 | 121 | 223 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 1.7E-21 | 121 | 223 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 3.0E-33 | 288 | 371 | IPR010525 | Auxin response factor |
| PROSITE profile | PS51745 | 20.735 | 616 | 696 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007389 | Biological Process | pattern specification process | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0048829 | Biological Process | root cap development | ||||
| GO:0051301 | Biological Process | cell division | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 735 aa Download sequence Send to blast |
MITFMDSKEK LKEGEKCLDP QLWHACAGGM VQMPLVNAKV FYFPQGHAEH ACGPVDFRNY 60 PRVPPYILCR VSAIKFMADP ETDEVYAKIR LVSLSSSEAG YEDNGIGGIN GADSQDKPAS 120 FAKTLTQSDA NNGGGFSVPR YCAETIFPRL DYSADPPVQT ILAKDVHGET WKFRHIYRGT 180 PRRHLLTTGW STFVNHKKLV AGDSIVFLRA ENGDLCVGIR RAKRGIGGGP ESSSGWNPAG 240 GNCAMPYGGF SSYLREDEGK LMRNGNGNAS NGSLMGKGKV GPESVLEAAT LASNGQPFEV 300 VYYPRASTPE FCVKASLVKA ALQIRWCPGM RFKMAFETED SSRISWFMGT ISSVQVSDPM 360 RWPESPWRLL QVSWDEPDLL QNVKRVSPWL VELVSNMPAI HLTPFSPPRK KMRLPQHPDF 420 PFEGQLPMPT FSGNHHLLGT SSPFGCLPDK TPAGMQGARH AHYGLSLSDI HLNKLQSGLF 480 PAGFPPLDHV ATPTKFSNNT MIQRPTMSEN VSCLLTMSHS PQSTKKSEDV KPPQLMLFGK 540 PILTEQQISL SSSGDTVSPV VTGNSLSDGS GDKMANHSEN SGSALHQQSV QERSSGEGFF 600 HWYKDTRHEA EANLETGHCK VFMESEDVGR TLDFSQFKSY DELYTKLADM FGIENSETLN 660 HVLYRDATGA VKHIGDEPFS DFVKTARRLT ILMDSGSNNV GICGRSRTYT DLIYELPIQI 720 IPVSWSIFLS RRRNT |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldu_A | 5e-81 | 3 | 392 | 32 | 388 | Auxin response factor 5 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
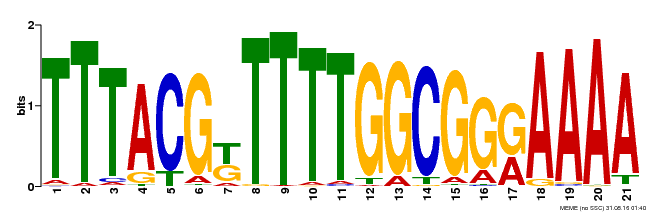
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00461 | DAP | Transfer from AT4G30080 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004304436.1 | 0.0 | PREDICTED: auxin response factor 18-like isoform X2 | ||||
| Swissprot | Q653H7 | 0.0 | ARFR_ORYSJ; Auxin response factor 18 | ||||
| TrEMBL | A0A2P6RAT8 | 0.0 | A0A2P6RAT8_ROSCH; Auxin response factor | ||||
| STRING | XP_004304436.1 | 0.0 | (Fragaria vesca) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF637 | 34 | 138 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G30080.1 | 0.0 | auxin response factor 16 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



