 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | FANhyb_rscf00000452.1.g00006.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Rosoideae; Potentilleae; Fragariinae; Fragaria
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 865aa MW: 94066.6 Da PI: 6.4714 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 66.6 | 3.4e-21 | 111 | 166 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+++l+L++rqVk+WFqNrR+++k
FANhyb_rscf00000452.1.g00006.1 111 KKRYHRHTPQQIQELEALFKECPHPDEKQRLELSRRLNLETRQVKFWFQNRRTQMK 166
688999***********************************************999 PP
| |||||||
| 2 | START | 206.7 | 9.3e-65 | 315 | 537 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE..EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT CS
START 1 elaeeaaqelvkkalaeepgWvkss..esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqW 72
ela++a++elvk+a+ +ep+W+++ e +n +e++++f++ + + +ea+r++g+v+ ++ lve+l+d++ +W
FANhyb_rscf00000452.1.g00006.1 315 ELALAAMDELVKLAQTDEPLWSLEGgrEVLNHEEYMRSFTPCIGlkpngFVTEASRETGMVIINSLALVETLMDSN-RW 392
5899********************999*************999899999***************************.** PP
-TT-S....EEEEEEEECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-T CS
START 73 detla....kaetlevissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppess 140
e+++ + +t++vissg galqlm aelq+lsplvp R++ f+R+++q+ +g+w++vdvSvd ++++ +
FANhyb_rscf00000452.1.g00006.1 393 LEMFPcmiaRTSTTDVISSGmggtrnGALQLMHAELQVLSPLVPvREVNFLRFCKQHAEGVWAVVDVSVDTIRDNSGAP 471
****************************************************************************9** PP
TSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 141 svvRaellpSgiliepksnghskvtwvehvdlkgrlphwllrslvksglaegaktwvatlqrqcek 206
++ +++lpSg+++++++ng+skvtwveh++++++++h l+r+l++sg+ +ga++wvatlqrqc++
FANhyb_rscf00000452.1.g00006.1 472 TFANCRRLPSGCVVQDMPNGYSKVTWVEHAEYDESQVHHLYRPLLSSGMGFGAQRWVATLQRQCQC 537
****************************************************************85 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.3E-21 | 97 | 168 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.4E-22 | 97 | 168 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 18.058 | 108 | 168 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 3.6E-19 | 109 | 172 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.29E-19 | 111 | 168 | No hit | No description |
| Pfam | PF00046 | 8.6E-19 | 111 | 166 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 143 | 166 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 43.593 | 306 | 540 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 3.3E-33 | 310 | 537 | No hit | No description |
| CDD | cd08875 | 4.29E-124 | 312 | 536 | No hit | No description |
| SMART | SM00234 | 8.3E-48 | 315 | 537 | IPR002913 | START domain |
| Pfam | PF01852 | 5.5E-57 | 315 | 537 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 2.51E-18 | 558 | 669 | No hit | No description |
| SuperFamily | SSF55961 | 2.51E-18 | 723 | 792 | No hit | No description |
| SuperFamily | SSF55961 | 2.51E-18 | 819 | 851 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 865 aa Download sequence Send to blast |
MSFGGFLDNS TGSSGGARIV ADIPYNHHPH HNANHTSMPS SAIAQPRLQT NADGGGDAAR 60 MAENFEGNNN VGGRRSREEE NEISRSGSDN MDGAGSGDEG DAADNSNPRK KKRYHRHTPQ 120 QIQELEALFK ECPHPDEKQR LELSRRLNLE TRQVKFWFQN RRTQMKTQLE RHENSLLRQE 180 NDKLRAENMS IRDAMRNPIC TNCGGPAMIG DISIEEQHLR IDNARLKDEL DRVCALAGKF 240 LGRPISSLGP SMGPPLPSSA LELGVGNNGF GGMSSVSTTM PLGPDFGAGL GGGMPLVAHT 300 RPVAGGLDER TMFLELALAA MDELVKLAQT DEPLWSLEGG REVLNHEEYM RSFTPCIGLK 360 PNGFVTEASR ETGMVIINSL ALVETLMDSN RWLEMFPCMI ARTSTTDVIS SGMGGTRNGA 420 LQLMHAELQV LSPLVPVREV NFLRFCKQHA EGVWAVVDVS VDTIRDNSGA PTFANCRRLP 480 SGCVVQDMPN GYSKVTWVEH AEYDESQVHH LYRPLLSSGM GFGAQRWVAT LQRQCQCLAI 540 LMSSTVPARD HANTITQSGR KSMLKLAQRM TDNFCAGVCA STVHKWNKLN AGNVDEDVRY 600 MTRESMDDPG EPPGIVLSAA TSVWLPVSPQ RLFNFLRDER LRSEWDILSN GGPMQEMAHI 660 AKGQDQGNCV SLLRARVSFL NFLSLNYLVQ YDACVPGRES EVKMAKRDGV FVWLCVGIMG 720 KVKVKWGEKG QAMNANQNSM LILQETCIDA AGSLVVYAPV DIPAMHVVMN GGDSAYVALL 780 PSGFAIVPDG PGSRGPGGAE GKAGQGGSNG NGGEARVSGS LLTMTFQILV NSLPSAKLTV 840 ESVETVNNLI SCTVQKIKGA LQCES |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
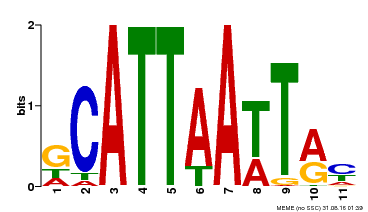
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004306832.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ANTHOCYANINLESS 2 isoform X1 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A2P6SGK8 | 0.0 | A0A2P6SGK8_ROSCH; Putative transcription factor & lipid binding Homobox-WOX family | ||||
| STRING | XP_004306832.1 | 0.0 | (Fragaria vesca) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1192 | 34 | 106 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



