 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | FANhyb_rscf00000253.1.g00010.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Rosoideae; Potentilleae; Fragariinae; Fragaria
|
||||||||
| Family | BBR-BPC | ||||||||
| Protein Properties | Length: 702aa MW: 77344.9 Da PI: 7.5135 | ||||||||
| Description | BBR-BPC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GAGA_bind | 389.4 | 5.3e-119 | 362 | 702 | 1 | 301 |
GAGA_bind 1 mdddgsre..rnkg.yye...........pa.aslkenlglqlmssiaerdakirernlalsekkaavaerdmaflqrd 64
mdd g++e r k+ +y+ p+ +s+k q+m+++aerda+i+ern+a+sekkaa+aerdmaflqrd
FANhyb_rscf00000253.1.g00010.1 362 MDDGGHHEngRLKAeQYKaaqgqwlmqhqPHqPSMK-----QIMALMAERDAAIQERNVAFSEKKAALAERDMAFLQRD 435
899999888999999999888877765542234555.....************************************** PP
GAGA_bind 65 kalaernkalverdnkllalllvensla.....salpvgvqvlsgtksidslqq.lse.pqledsavelreeeklealp 136
+a+aern+a++erdn++++l+++ens+ s++p+g+q+++g k++++ qq +++ p++++ ++++re++++++lp
FANhyb_rscf00000253.1.g00010.1 436 AAIAERNNAIMERDNAIATLQYRENSSLtsgnmSSCPPGCQISRGIKHMHNPQQhVHHlPHMNEASYGTREMHTNDSLP 514
*************************988899999********************55559******************** PP
GAGA_bind 137 ieeaaeeakekkkkkkrqrakkpkekkakkkkkksekskkkvkkesader............................. 186
+ ++ a ++++ +k+pke+k+ ++kk++ks +kvk+es+d +
FANhyb_rscf00000253.1.g00010.1 515 LPPDPPPAPKSRQ------TKRPKEPKTIPPNKKPAKSPRKVKRESEDLNkmtfdrlhewkggqemcggaedlnkqlgv 587
9887766555544......4577777777777777778888888887744478888999******************** PP
GAGA_bind 187 skaekksidlvlngvslDestlPvPvCsCtGalrqCYkWGnGGWqSaCCtttiSvyPLPvstkrrgaRiagrKmSqgaf 265
sk+++k++dl+ln+v++D+st+P+PvCsCtG+lrqCYkWGnGGWqS+CCttt+S+yPLP+++++r+aR++grKmS++af
FANhyb_rscf00000253.1.g00010.1 588 SKSDWKCQDLGLNQVAYDDSTMPAPVCSCTGVLRQCYKWGNGGWQSSCCTTTMSIYPLPAVPNKRHARVGGRKMSGSAF 666
******************************************************************************* PP
GAGA_bind 266 kklLekLaaeGydlsnpvDLkdhWAkHGtnkfvtir 301
+klL++LaaeG+dlsnpvDLkd WAkHGtn+++ti+
FANhyb_rscf00000253.1.g00010.1 667 NKLLSRLAAEGHDLSNPVDLKDNWAKHGTNRYITIK 702
***********************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00645 | 5.2E-97 | 4 | 209 | IPR000668 | Peptidase C1A, papain C-terminal |
| PRINTS | PR00705 | 2.5E-6 | 11 | 26 | IPR000668 | Peptidase C1A, papain C-terminal |
| SuperFamily | SSF54001 | 1.42E-77 | 14 | 210 | No hit | No description |
| CDD | cd02248 | 6.91E-91 | 15 | 208 | No hit | No description |
| Pfam | PF00112 | 3.5E-71 | 15 | 208 | IPR000668 | Peptidase C1A, papain C-terminal |
| Gene3D | G3DSA:3.90.70.10 | 3.3E-81 | 15 | 209 | No hit | No description |
| PROSITE pattern | PS00639 | 0 | 152 | 162 | IPR025660 | Cysteine peptidase, histidine active site |
| PRINTS | PR00705 | 2.5E-6 | 154 | 164 | IPR000668 | Peptidase C1A, papain C-terminal |
| PROSITE pattern | PS00640 | 0 | 169 | 188 | IPR025661 | Cysteine peptidase, asparagine active site |
| PRINTS | PR00705 | 2.5E-6 | 169 | 175 | IPR000668 | Peptidase C1A, papain C-terminal |
| SuperFamily | SSF57277 | 1.69E-6 | 226 | 259 | No hit | No description |
| SMART | SM00277 | 3.8E-22 | 229 | 286 | IPR000118 | Granulin |
| Pfam | PF00396 | 2.2E-10 | 240 | 287 | IPR000118 | Granulin |
| Pfam | PF06217 | 1.0E-102 | 362 | 702 | IPR010409 | GAGA-binding transcriptional activator |
| SMART | SM01226 | 1.1E-172 | 362 | 702 | IPR010409 | GAGA-binding transcriptional activator |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0006508 | Biological Process | proteolysis | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008234 | Molecular Function | cysteine-type peptidase activity | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 702 aa Download sequence Send to blast |
MYSLEYINIF NICTGACWSF SATGAIEGIN KIVTGSLVSL SEQELIDCDR VYPNSGCNGG 60 LMDDAFQFII DNNGIDTEED YPYQGADGTC NKQKLKRHVV TIDGYTDVPA NNEEQLLKAV 120 ATQPVSVGIA GSGREFQFYS KGIFAGPCST TLDHAVLIVG YGSENGVDYW IVKNSWGKNW 180 GMNGYIHMLR DHSNSKGLCG INMLASYPTK TGENPPFPPP SPPPGPTKCD LFSKCGVGET 240 CCCAKKILGI CLSWRCCELA SAVCCKDRLH CCPHDYPICD TERNYCLQPK SLKILLLLPP 300 DHHSFTAFPS SSSSSSQIRV SPSYLGLVFG AGWDWFGVLG FMGSGYLILG SELDSVLCSE 360 LMDDGGHHEN GRLKAEQYKA AQGQWLMQHQ PHQPSMKQIM ALMAERDAAI QERNVAFSEK 420 KAALAERDMA FLQRDAAIAE RNNAIMERDN AIATLQYREN SSLTSGNMSS CPPGCQISRG 480 IKHMHNPQQH VHHLPHMNEA SYGTREMHTN DSLPLPPDPP PAPKSRQTKR PKEPKTIPPN 540 KKPAKSPRKV KRESEDLNKM TFDRLHEWKG GQEMCGGAED LNKQLGVSKS DWKCQDLGLN 600 QVAYDDSTMP APVCSCTGVL RQCYKWGNGG WQSSCCTTTM SIYPLPAVPN KRHARVGGRK 660 MSGSAFNKLL SRLAAEGHDL SNPVDLKDNW AKHGTNRYIT IK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1aec_A | 4e-75 | 15 | 210 | 23 | 217 | ACTINIDIN |
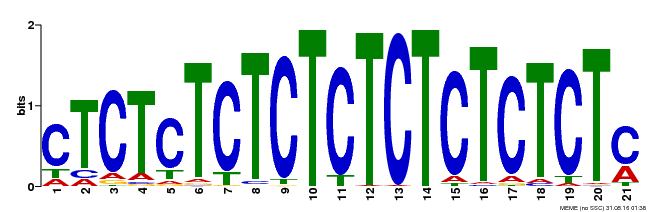
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00540 | DAP | Transfer from AT5G42520 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004291398.1 | 0.0 | PREDICTED: protein BASIC PENTACYSTEINE6 | ||||
| STRING | XP_004291398.1 | 0.0 | (Fragaria vesca) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF6905 | 34 | 49 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G42520.1 | 1e-147 | basic pentacysteine 6 | ||||



