 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 462919188 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Eragrostideae; Eragrostidinae; Eragrostis
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 1094aa MW: 121004 Da PI: 7.4466 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 90.4 | 1.4e-28 | 497 | 548 | 1 | 52 |
RWP-RK 1 aekeisledlskyFslpikdAAkeLgvclTvLKriCRqyGIkRWPhRkiksl 52
aek+isl++l++yFs+++k+AAk+Lgvc+T++KriCRq+GI+RWP+Rki+++
462919188 497 AEKTISLDVLQQYFSGSLKNAAKSLGVCPTTMKRICRQHGISRWPSRKINKV 548
589***********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51519 | 17.541 | 487 | 568 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 7.8E-26 | 500 | 548 | IPR003035 | RWP-RK domain |
| SuperFamily | SSF54277 | 4.25E-20 | 767 | 862 | No hit | No description |
| SMART | SM00666 | 5.7E-24 | 777 | 859 | IPR000270 | PB1 domain |
| Pfam | PF00564 | 7.1E-15 | 777 | 858 | IPR000270 | PB1 domain |
| PROSITE profile | PS51745 | 18.652 | 777 | 859 | IPR000270 | PB1 domain |
| Gene3D | G3DSA:3.10.20.240 | 1.6E-24 | 777 | 858 | No hit | No description |
| CDD | cd06407 | 1.10E-32 | 778 | 858 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1094 aa Download sequence Send to blast |
MGNSDTTSKR VEQINHKWQL HLSLDGDGTD NSSLFKERLT QALRYFKDST DQHLLVQVWA 60 PVKNGDRYVL TTSGQPFVLD HQSIGLLQYR AVSMMYMFSV DGDNAGELGL PGRVYKQKVP 120 EWTPNVQYYS SNEYPRLNHA ISYNVHGTVA LPVFDPSAQS CIAVVELIMT SKKINYACEV 180 DKVCKALEAV NLKSTEIFDH PNVQICNEGR QTALVEILEI LTVVCEEHKL PLAQTWVPCK 240 YRSVLAHGGG LKKSCLSFDG SCMGEVCMST SDVAFHVIDA HMWGFRDACV EHHLQRGQGV 300 PGKAFISHKP CFSKDIRQFC KLEYPLVHYA RMFGLAGCLA ICLQSCYTGH DDYILEFFLP 360 PDCIDEDDQN ALLESILTRM KKCLRSLKVV GDRDLNGVSL QLGNVLKIEN EEFKKDVQFD 420 NSEGCLRESP EGDTRGRVHE FDTETKRVSN MPEGHILADD QSQDNGTSAT RQNGSGASDS 480 SLLHKTNKPP ERRRGKAEKT ISLDVLQQYF SGSLKNAAKS LGVCPTTMKR ICRQHGISRW 540 PSRKINKVNR SLSKLKQVIE SVQGSDAAFN LTSITGPLPI PVGPSSDSLN IEKVTQSKVA 600 DLSNPADGDR DSSLQKSLEN DGHFGILMAQ QGFMDNNNSK NHLKMMVILQ GFMDNNNDAQ 660 LEADKASHSR SSSGEGSINS RTSEGSCQGS PANQTFVCKP IASTFAEPQI NPEEFTKEPF 720 QEPQLPLSRM LIEDSGSSKD LKNLFTSAAD QPFFATPTDQ PFFAPPSNLR SMKHSGTVTI 780 KASFKEDIVR FRFPCSGSVT VLKDEVAKRL RMDVGMFDIK YLDDDHEWVK LACNADLEEC 840 MEISRHSGTH VIRLLVSDIA AHLGSSCGSS EKTSAGFGLG GVVFSLKHPT IPSDSVGPIV 900 LICTLEHIGF MRGTNRESRE GVLFVRNGFD TVIPSREFGQ PNIPVNKWPY AKHYPEFANF 960 ATFGSRQDKE ANWRTSSWKD EWLVFSTMIG GSKTCVSVTA AAFQHGGPAS VKSGESTAIG 1020 EPKYQIPFGI PKGEAVKTPH LIFKNRNISS ISENTLHFKG TRVNRLGKRN MGVQFNLVCA 1080 ATNTKVKLGS INKS |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
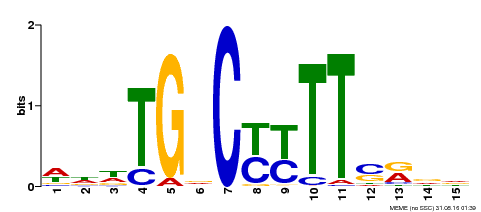
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 462919188 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT085359 | 0.0 | BT085359.2 Zea mays full-length cDNA clone ZM_BFc0002J20 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025814939.1 | 0.0 | protein NLP3 isoform X1 | ||||
| Swissprot | Q5NB82 | 0.0 | NLP3_ORYSJ; Protein NLP3 | ||||
| TrEMBL | A0A2S3HZV8 | 0.0 | A0A2S3HZV8_9POAL; Uncharacterized protein | ||||
| STRING | Si000216m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4778 | 36 | 60 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G64530.1 | 0.0 | Nin-like family protein | ||||



