 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 462913299 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Eragrostideae; Eragrostidinae; Eragrostis
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 474aa MW: 51666 Da PI: 9.1884 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 130 | 8.4e-41 | 51 | 127 | 2 | 78 |
-SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 2 CqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
C vegC+adls++++yhrrhkvCe+hsk+pvv+v+g++qrfCqqCsrfh l efDe krsCr+rL++hn+rrrk+q+
462913299 51 CSVEGCAADLSKCRDYHRRHKVCEAHSKTPVVTVAGQQQRFCQQCSRFHLLCEFDEVKRSCRKRLDGHNRRRRKPQP 127
**************************************************************************987 PP
| |||||||
| 2 | SBP | 46.2 | 1.2e-14 | 224 | 259 | 43 | 78 |
-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 43 CqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
C fh l efDe krsCr+rL++hn+rrrk+q+
462913299 224 LCPCREFHLLCEFDEVKRSCRKRLDGHNRRRRKPQP 259
5569*****************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 5.7E-33 | 43 | 112 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.129 | 48 | 125 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.03E-38 | 49 | 130 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 3.3E-29 | 51 | 124 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 13.084 | 180 | 257 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 2.35E-12 | 223 | 262 | IPR004333 | Transcription factor, SBP-box |
| Gene3D | G3DSA:4.10.1100.10 | 8.3E-4 | 226 | 244 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 5.1E-7 | 227 | 256 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048653 | Biological Process | anther development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 474 aa Download sequence Send to blast |
MPRCLLGTSA RWSAPAAAAP WTSSSAGRSP SSSAPAKRPR AGQAQQAVPA CSVEGCAADL 60 SKCRDYHRRH KVCEAHSKTP VVTVAGQQQR FCQQCSRFHL LCEFDEVKRS CRKRLDGHNR 120 RRRKPQPDPM NPGALFANHH GMTRFTSYPQ LFSAPSMADS KWPATTIVKA ETDAFQDHYY 180 PAVHLNNGAV SLFNGKDRKH FPFLTPHGGI TRTTDSVLFS DAFLCPCREF HLLCEFDEVK 240 RSCRKRLDGH NRRRRKPQPD PMNPGALFAN HHGMTRFTSY PQLFSAPSMA DSKWPATIVK 300 AETDAFQDHY YPAVHLNNGA VSLFNGKDRK HFPFLTPHGG DAAAALGCQP FTITPSSESN 360 SKQSNGNCAL SLLSDNSTPA QLMIPTGQPL GAALHYGNVA RLPDDGDVSL TGMSYVSMGD 420 KQTSIVATSA GHTATAASPA PATQLQYHGY YHVNGGDQGN SDGASIQALP FSSW |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 8e-26 | 48 | 124 | 8 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 239 | 256 | KRSCRKRLDGHNRRRRKP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' (By similarity). May be involved in panicle development. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00555 | DAP | Transfer from AT5G50570 | Download |
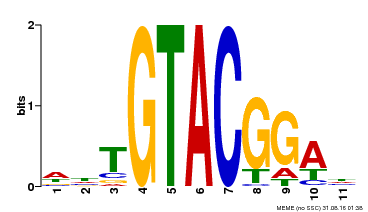
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 462913299 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Negatively regulated by microRNAs miR156b and miR156h. {ECO:0000305|PubMed:16861571}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004971091.1 | 1e-117 | squamosa promoter-binding-like protein 2 | ||||
| Swissprot | Q0JGI1 | 1e-78 | SPL2_ORYSJ; Squamosa promoter-binding-like protein 2 | ||||
| TrEMBL | K3XIT1 | 1e-116 | K3XIT1_SETIT; Uncharacterized protein | ||||
| STRING | Si001804m | 1e-117 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP10456 | 30 | 38 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G50670.1 | 2e-31 | SBP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



