 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 462909130 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Eragrostideae; Eragrostidinae; Eragrostis
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 452aa MW: 49508.2 Da PI: 6.27 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 101.9 | 4e-32 | 266 | 320 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
k+rlrWt eLHerFveav++L G+ekAtPk +l+lmkv+gLt++hvkSHLQkYRl
462909130 266 KTRLRWTLELHERFVEAVNKLDGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRL 320
68****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 11.574 | 263 | 323 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 4.12E-17 | 263 | 320 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 3.1E-30 | 264 | 321 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.3E-24 | 266 | 321 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 3.6E-8 | 268 | 319 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 1.5E-21 | 356 | 402 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 452 aa Download sequence Send to blast |
MSSQSSVTVK QMAAPDKTVH ASTSTQPLVH KLLDAKLDHQ VLVDDTLSST SQSSSIKTEL 60 MRSSSFSMSL TLSLQKRSPD TDPESPLSHV SHPNFSDPIP SNSSTFCTSL FSSSLSNPEP 120 TRQIGTLPFL PHPPKCEQQV SAGQSSSSSL LLSGDISNSI DESEHSDDLK DLLNLSGDGS 180 DGSYHGENNA LAFSEHMEFQ FLSEQLGIAI TDNESPRLDD IYGTPPQLSS LPVLSCSNQS 240 IQNPGSPVKV HLNSPRSSSG SATTNKTRLR WTLELHERFV EAVNKLDGPE KATPKGVLKL 300 MKVEGLTIYH VKSHLQKYRL AKYLPEPRED KKAPSEDKKA QSDSSSNDSG KKKNLQVAEA 360 LRMQMEVQKQ LHEQLEVQRQ LQLRIEEHAR YLQKLLEEQQ KAGNSLSLKT PTEAQAESPE 420 LTSKERGHEH LQSTLSGSML DEATGIALAP AK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 2e-25 | 266 | 323 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 2e-25 | 266 | 323 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 2e-25 | 266 | 323 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 2e-25 | 266 | 323 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 2e-25 | 266 | 323 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 2e-25 | 266 | 323 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 2e-25 | 266 | 323 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 2e-25 | 266 | 323 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in phosphate starvation signaling (PubMed:26082401). Binds to P1BS, an imperfect palindromic sequence 5'-GNATATNC-3', to promote the expression of inorganic phosphate (Pi) starvation-responsive genes (PubMed:26082401). Functionally redundant with PHR1 and PHR2 in regulating Pi starvation response and Pi homeostasis (PubMed:26082401). {ECO:0000269|PubMed:26082401}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00354 | DAP | Transfer from AT3G13040 | Download |
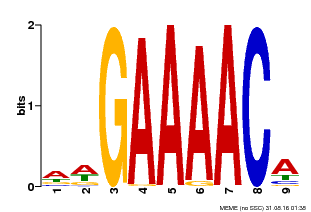
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 462909130 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated under Pi starvation conditions. {ECO:0000269|PubMed:26082401}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025812859.1 | 0.0 | protein PHOSPHATE STARVATION RESPONSE 3 | ||||
| Refseq | XP_025812860.1 | 0.0 | protein PHOSPHATE STARVATION RESPONSE 3 | ||||
| Refseq | XP_025812862.1 | 0.0 | protein PHOSPHATE STARVATION RESPONSE 3 | ||||
| Refseq | XP_025812863.1 | 0.0 | protein PHOSPHATE STARVATION RESPONSE 3 | ||||
| Swissprot | Q6YXZ4 | 1e-160 | PHR3_ORYSJ; Protein PHOSPHATE STARVATION RESPONSE 3 | ||||
| TrEMBL | A0A3L6PFL7 | 0.0 | A0A3L6PFL7_PANMI; Uncharacterized protein | ||||
| STRING | Pavir.J27649.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4526 | 37 | 69 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G13040.2 | 2e-71 | G2-like family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



