 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thhalv10008152m | ||||||||
| Common Name | EUTSA_v10008152mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Eutremeae; Eutrema
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 335aa MW: 37581.9 Da PI: 5.7058 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 56.7 | 5.4e-18 | 15 | 62 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd++l+++ +++G g W+t +++ g++R++k+c++rw +yl
Thhalv10008152m 15 KGAWTPEEDQKLICYLNRHGEGGWRTLPEKAGLKRCGKSCRLRWANYL 62
79********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 52.7 | 9.5e-17 | 68 | 112 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
rg +T++E+ ++ +++++G++ W++Iar ++ gRt++++k++w+++
Thhalv10008152m 68 RGEFTEDEERSIISLHALHGNK-WSAIARGLP-GRTDNEIKNYWNTH 112
899*******************.*********.************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 15.229 | 10 | 62 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 4.67E-29 | 12 | 109 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.6E-13 | 14 | 64 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.9E-16 | 15 | 62 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 8.0E-23 | 16 | 69 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 6.01E-10 | 17 | 62 | No hit | No description |
| PROSITE profile | PS51294 | 25.381 | 63 | 117 | IPR017930 | Myb domain |
| SMART | SM00717 | 3.4E-16 | 67 | 115 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.0E-15 | 68 | 112 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 6.1E-26 | 70 | 118 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.15E-11 | 70 | 113 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009625 | Biological Process | response to insect | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009682 | Biological Process | induced systemic resistance | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0009759 | Biological Process | indole glucosinolate biosynthetic process | ||||
| GO:0052544 | Biological Process | defense response by callose deposition in cell wall | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 335 aa Download sequence Send to blast |
MVRTPCCKAE LGLKKGAWTP EEDQKLICYL NRHGEGGWRT LPEKAGLKRC GKSCRLRWAN 60 YLRPDIKRGE FTEDEERSII SLHALHGNKW SAIARGLPGR TDNEIKNYWN THIKKRLIRK 120 GVDPVTQNSL ISGRGKSENP PGFPQKQKVI QTITSDDDLD DEKVNNNNKK SGVSSARFLN 180 RVANRFGKII NHSVLSGIIG SGGPLTTTTT STTSVTVDSE SDKSTSSSFT PTSNLLCQMP 240 VHDNGNVTSS SSTFSDSSVN DPLLMYCDNK DNMGFSRFLS DEDFMMLEES SIENTEFMKE 300 LTRFLEEDEN DVVDEVTPVY EYQDIFQEID HYFA* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 3e-28 | 13 | 117 | 5 | 108 | B-MYB |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor positively regulating indolic glucosinolate biosynthetic pathway genes. {ECO:0000269|PubMed:17461791, ECO:0000269|PubMed:23580754, ECO:0000269|PubMed:23943862}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
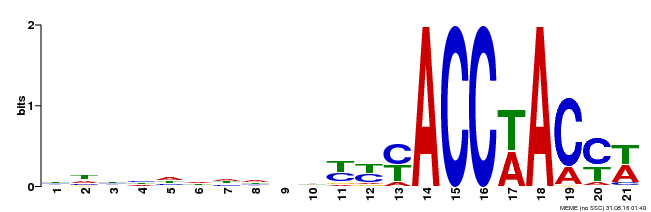
| Motif ID | Method | Source | Motif file |
| MP00147 | DAP | Transfer from AT1G18570 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Thhalv10008152m |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by touch or wounding. {ECO:0000269|PubMed:17461791, ECO:0000269|PubMed:23115560}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006416592.1 | 0.0 | transcription factor MYB51 | ||||
| Swissprot | O49782 | 1e-159 | MYB51_ARATH; Transcription factor MYB51 | ||||
| TrEMBL | V4KB56 | 0.0 | V4KB56_EUTSA; Uncharacterized protein | ||||
| STRING | XP_006416592.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4 | 28 | 2646 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G18570.1 | 1e-127 | myb domain protein 51 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thhalv10008152m |
| Entrez Gene | 18993054 |



