 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thhalv10003564m | ||||||||
| Common Name | EUTSA_v10003564mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Eutremeae; Eutrema
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1064aa MW: 119083 Da PI: 6.3242 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 187.1 | 1.8e-58 | 19 | 136 | 2 | 118 |
CG-1 2 lke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptf 94
l+e ++rwl++ ei++iL n++k+++++e+++rp+sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+hekLKvg ++vl+cyYah+e+n++f
Thhalv10003564m 19 LSEaQHRWLRPAEICEILRNYQKFHIASEPPNRPPSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHEKLKVGRIDVLHCYYAHGEDNENF 112
45559***************************************************************************************** PP
CG-1 95 qrrcywlLeeelekivlvhylevk 118
qrrcyw+Le++l +iv+vhylevk
Thhalv10003564m 113 QRRCYWMLEQDLMHIVFVHYLEVK 136
*********************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 85.888 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 2.5E-85 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 7.7E-52 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| SuperFamily | SSF81296 | 1.52E-14 | 470 | 556 | IPR014756 | Immunoglobulin E-set |
| SuperFamily | SSF48403 | 3.58E-18 | 652 | 764 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 18.643 | 653 | 764 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 1.0E-6 | 654 | 730 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.5E-17 | 654 | 765 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 6.95E-14 | 659 | 762 | No hit | No description |
| PROSITE profile | PS50088 | 10.633 | 703 | 735 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.0046 | 703 | 732 | IPR002110 | Ankyrin repeat |
| SMART | SM00015 | 0.32 | 878 | 900 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF52540 | 1.06E-7 | 878 | 928 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| PROSITE profile | PS50096 | 7.858 | 880 | 908 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0021 | 880 | 899 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0092 | 901 | 923 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.304 | 902 | 926 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 4.7E-4 | 904 | 923 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1064 aa Download sequence Send to blast |
MADRGSFGFA PRLDIEQLLS EAQHRWLRPA EICEILRNYQ KFHIASEPPN RPPSGSLFLF 60 DRKVLRYFRK DGHNWRKKKD GKTVKEAHEK LKVGRIDVLH CYYAHGEDNE NFQRRCYWML 120 EQDLMHIVFV HYLEVKGNRM SSSGIKENNS NSLSGTTSVN IDSIATPSST LSPLCEDADS 180 GDSRQANSSL QPNPELQTVA PQIRHQQNAR TINSYNPTSI LGNRHGWTSA PGIGIVSQVH 240 GNRVKESDSQ RSVDVPTWDA SFENSLARYQ NLPYNAPLTQ TQPFNAGLMP VEGNKEKGSL 300 LTAEHLRSPL QNQVNWQIPV QDSLPVQKWP MDSHSGMTDS TDLALLGQRA HENCGTFSSL 360 LGSQNQQPVG GSFQAPFTSI EAAYIPKLGP EDLLYEASAN QTLPLRKSLL KEEDSLKKVD 420 SFSRWVSNEL GEMEDLQMQS SSGGIPWTTV ETAAAASSLS PSLSEDQRFT IIDFWPKWTQ 480 TDSEVEVMVI GTFLLTPHEV TSYNWSCMFG EVEVPAEILV DGVLCCHAPP HEVGQVPFYI 540 TCSDRFSCSE VREFDFLPGS TKKLNTSDIY GAFTNEASLH LRFENLLARI SSVQKHHIFE 600 NVGVKRTKIS RMMLLKDEKE SLLPGTIQKD LPEPEATERL IREEFEDRLY IWLIHKVTEE 660 GKGPNILDEE GQGVLHLVAA LGYDWAIKPI LAAGVSINFR DANGWSALHW AAFCGREDTV 720 AVLVSLGADA GALTDPSPEL PLGKTAADLA YGNGHRGISG FLAESSLTSY LEKLTVDAKE 780 NSSADSSRVK AVQTVAERTA TPMSYGDVPE TLSMKDSLTA VFNATQAADR LHQVFRMQSF 840 QRKQLSEFGY NSEFDISDEL AVSFAAAKTK KPGHSNGAVH AAAVQIQKKY RGWKKRKEFL 900 LIRQRIVKIQ AHVRGHQVRK QYRAIIWSVG LLEKIILRWR RKGSGLRGFK RDTITKPPEP 960 VSAAPPPQED DYDFLKEGRK QTEERLQKAL TRVKSMAQYP EARAQYRRLL TVVEGFRENE 1020 ASSSSAMNNN NNTEEAANYN EEDDLIDIDS LLDNDTFMSL AFE* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
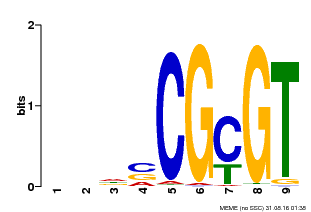
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Thhalv10003564m |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK229403 | 0.0 | AK229403.1 Arabidopsis thaliana mRNA for Calmodulin-binding transcription activator 2, complete cds, clone: RAFL16-66-C08. | |||
| GenBank | BT010874 | 0.0 | BT010874.1 Arabidopsis thaliana At5g64220 gene, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006394184.1 | 0.0 | calmodulin-binding transcription activator 2 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | V4K0S6 | 0.0 | V4K0S6_EUTSA; Uncharacterized protein | ||||
| STRING | XP_006394184.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4082 | 24 | 52 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thhalv10003564m |
| Entrez Gene | 18012634 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



