 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thhalv10001295m | ||||||||
| Common Name | EUTSA_v10001295mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Eutremeae; Eutrema
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 936aa MW: 104266 Da PI: 5.8027 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 135.3 | 1.9e-42 | 144 | 221 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+Ceadls++k+yhrrhkvCe+hska++++v g++qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
Thhalv10001295m 144 ACQVENCEADLSKVKDYHRRHKVCEMHSKATSAIVGGIMQRFCQQCSRFHVLQEFDEGKRSCRRRLAGHNKRRRKTNP 221
5**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 3.0E-34 | 138 | 206 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.633 | 142 | 219 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 2.22E-39 | 144 | 223 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 7.4E-31 | 145 | 218 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF48403 | 3.58E-8 | 725 | 826 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 2.47E-9 | 727 | 823 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 6.4E-8 | 727 | 824 | IPR020683 | Ankyrin repeat-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 936 aa Download sequence Send to blast |
MEARIEGEAQ HLYGLRAMDL RSVGKRSVEW DLNDWKWDGD LFVATQLNPG GGETMGRQFF 60 PIGNSSNSSS SCSDEGNNTE LTVTRPREVE KKRRAVATGE DNNGGGGGGL TLKLGDDGYD 120 LNGEREGGSA AKKTKFGGGT NRAACQVENC EADLSKVKDY HRRHKVCEMH SKATSAIVGG 180 IMQRFCQQCS RFHVLQEFDE GKRSCRRRLA GHNKRRRKTN PEPGANGNPL SDDHQSSNFL 240 LISLLKILSN MHSNRSDHPG DQDLMSHLLK SLVSHAGEQL GKNLVELLLQ GGSQASLNIG 300 NSNLLAIEQA PQEDLKQAPE IPRQELYANG TATENRSEKQ VKVNDFDLND IYIDSDDGTD 360 IERSPPPTNP ATSSLDYPSW IHQSSPPQTS RNSDSASDQS PSSSSEDAQM RTGRIVFKLF 420 GKEPNDFPVV LRGQILDWLS HSPTDMESYI RPGCIVLTIY LRQAETAWED LSDDLGFSLR 480 KLLDLSDDPL WTTGWIYVRV QNQLAFVFDG QVVVDTSLPL RSRDYSHIMS VRPLAVATTG 540 KAQFTVKGIN LRRPGTRLLC AVEGKYLIQE TKHDSTKEND DCNENNEIVE CVSFSCDLPI 600 TSGRGFMEIE DQGLSSSFFP FIVVEEDDVC SEIRILETTL EFTETDSAKQ AMDFIHEIGW 660 LLHRSKLGES DPNPDVFPLE RFKWLIEFSM DREWCAVIRK LLNMFFEGAV GDSSSSSDAA 720 LSELCLLHRA VRKNSKPMVE MLLRYVPKQQ RHSLFRPDAT GPAGLTPLHI AAGKDCSEDV 780 LDALTEDPAM VGIKAWKTSR DSTGFTPEDY ARLRGHFSYI HLIQRKINKK STTEDHVVVN 840 IPVSFSDREM KESKSGPVAS ALEITTQTNQ LQCKLCDHKL VYGTARRSVA YRPAMLSMVA 900 IAAVCVCVAL LFKSCPEVLY VFQPFRWELL DYGTR* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 8e-32 | 145 | 218 | 11 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00603 | PBM | Transfer from AT2G47070 | Download |
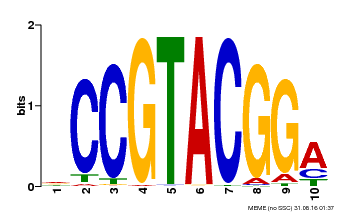
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Thhalv10001295m |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KC815948 | 0.0 | KC815948.1 Arabis alpina SQUAMOSA promoter binding protein-like protein 1 (SPL1) gene, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006397898.1 | 0.0 | squamosa promoter-binding-like protein 1 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | V4N2D6 | 0.0 | V4N2D6_EUTSA; Uncharacterized protein | ||||
| STRING | XP_006397898.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3030 | 27 | 64 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thhalv10001295m |
| Entrez Gene | 18015720 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



