 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010941149.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Arecoideae; Cocoseae; Elaeidinae; Elaeis
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 342aa MW: 36765.5 Da PI: 7.9549 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 42.4 | 1.7e-13 | 77 | 121 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT++E++++++a +++ + Wk+I + +g +t q++s+ qky
XP_010941149.1 77 RESWTEQEHDKFLEALQLFDRD-WKKIEAFVG-SKTVIQIRSHAQKY 121
789*****************77.*********.*************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.12E-16 | 71 | 127 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 15.408 | 72 | 126 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.2E-6 | 75 | 127 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.6E-18 | 75 | 124 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 5.3E-11 | 76 | 124 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.9E-11 | 77 | 121 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.20E-8 | 79 | 122 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 342 aa Download sequence Send to blast |
MVSVTAKPAD GGGGGGAAVL FFSDPHHYKQ PGSPMALPGL AGGGSLPTPS AAAATTASSS 60 DDASKKIRKP YTITKSRESW TEQEHDKFLE ALQLFDRDWK KIEAFVGSKT VIQIRSHAQK 120 YFLKVQKNGT SEHVPPPRPK RKAAHPYPQK ASKTAPIQAS SCLLEPGYAL TTDASSVLRN 180 STTNAIVSPR AHSSVQPVSA SHAIKDDVGP IGAVMANNCC CSSTESPSVT WQTCETTDQE 240 NHVPSLRVMP DFTQVYSFLG SVFEPNTSGH LLKLKEMDPI DVETVLLLMK NLSINLISPE 300 FEPHRRLLSS YSAGTEEVKP GSANNSPQVS ETMNTSFMVK GE |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00625 | PBM | Transfer from PK02532.1 | Download |
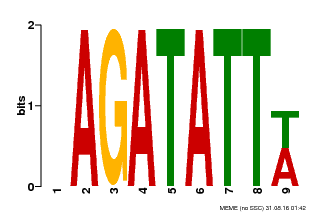
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_029116176.1 | 0.0 | protein REVEILLE 6 | ||||
| Swissprot | Q8H0W3 | 1e-101 | RVE6_ARATH; Protein REVEILLE 6 | ||||
| TrEMBL | A0A2H3Y8A3 | 0.0 | A0A2H3Y8A3_PHODC; protein REVEILLE 5-like isoform X6 | ||||
| STRING | XP_008794784.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1096 | 38 | 111 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G52660.1 | 5e-86 | MYB_related family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



