 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010939189.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Arecoideae; Cocoseae; Elaeidinae; Elaeis
|
||||||||
| Family | TCP | ||||||||
| Protein Properties | Length: 349aa MW: 39319.3 Da PI: 8.873 | ||||||||
| Description | TCP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCP | 94.1 | 2.8e-29 | 72 | 181 | 3 | 89 |
TCP 3 gkkdrhskihTkvggRdRRvRlsaecaarfFdLqdeLGfdkdsktieWLlqqakpaikeltgtssssaseceaesssssasnsssg......... 88
g+kdrhsk+ T +g+RdRRvRls++ a++++dLqd+LG+ ++sk ++WLl++a+++i++l+ ++ +++ + + s+s ++s++ ++
XP_010939189.1 72 GGKDRHSKVRTIKGLRDRRVRLSVPAAIQLYDLQDKLGLNQPSKVVDWLLNAAQHEIDKLPPLQLPPEFFI-QFSQSMATSHKLAPnkapsflpf 165
78*************************************************************88888555.33333333322222233445555 PP
TCP 89 ...............k 89
+
XP_010939189.1 166 rdgaelvyddrarqlA 181
5555555444444331 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51369 | 28.596 | 73 | 131 | IPR017887 | Transcription factor TCP subgroup |
| Pfam | PF03634 | 4.5E-28 | 73 | 203 | IPR005333 | Transcription factor, TCP |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0030154 | Biological Process | cell differentiation | ||||
| GO:0045962 | Biological Process | positive regulation of development, heterochronic | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 349 aa Download sequence Send to blast |
MMKSPPYAGN PQLQLTNCLQ SDESITMIRD SREEELLTKQ GGGTNDDRFA VSSRLWSALK 60 NPRIVRVSRT FGGKDRHSKV RTIKGLRDRR VRLSVPAAIQ LYDLQDKLGL NQPSKVVDWL 120 LNAAQHEIDK LPPLQLPPEF FIQFSQSMAT SHKLAPNKAP SFLPFRDGAE LVYDDRARQL 180 ASSSTMANGD MLDNNTGANE MMVQKSTFWS SNVLMKNKEK ATMKEPNIDK GSMIKGNDVG 240 SDGMQNNAMY TSYHHWNPPN EHISHLGSHL PQEEELHNYQ PGSMFAAYMN ASTDFAKQYH 300 HLDMANSASK FLPSTSLRNS LQFTGDPSLK LLQPNFGFKH HHPQDHQEG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5zkt_A | 5e-18 | 78 | 132 | 1 | 55 | Putative transcription factor PCF6 |
| 5zkt_B | 5e-18 | 78 | 132 | 1 | 55 | Putative transcription factor PCF6 |
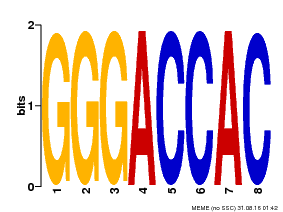
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00008 | PBM | Transfer from 496250 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010939189.1 | 0.0 | transcription factor TCP13 isoform X1 | ||||
| TrEMBL | A0A2H3XZZ9 | 0.0 | A0A2H3XZZ9_PHODC; transcription factor TCP13-like | ||||
| STRING | XP_008791353.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2490 | 35 | 84 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G60970.1 | 1e-42 | TEOSINTE BRANCHED 1, cycloidea and PCF transcription factor 5 | ||||



