 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010934336.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Arecoideae; Cocoseae; Elaeidinae; Elaeis
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 706aa MW: 76767.8 Da PI: 6.8475 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 73.8 | 2.1e-23 | 125 | 226 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSS CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrs 88
f k+lt+sd+++ g +++p+ +ae++ ++ + t+ +d +g++W++++iyr++++r++lt+GW+ Fv++++L +gD++vF + +
XP_010934336.1 125 FAKTLTQSDANNGGGFSVPRYCAETIfprldysADPPVQ--TVLAKDVHGEVWKFRHIYRGTPRRHLLTTGWSTFVNQKKLVAGDSIVFL--RAE 215
89****************************996555555..8999*********************************************..448 PP
EE..EEEEE-S CS
B3 89 efelvvkvfrk 99
++ l+v+++r+
XP_010934336.1 216 NGDLCVGIRRA 226
8999****997 PP
| |||||||
| 2 | Auxin_resp | 114.8 | 7.8e-38 | 295 | 378 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrl.sGtvvgvsdldpvrWpnSkWrsLk 83
aa+ a+++++Fe +Y+Pra+t+eF+vk++ v++a+++++++GmRfkmafetedss++++ +Gt+++v+ +dp+rWpnS+Wr+L+
XP_010934336.1 295 AATLAANGQPFEAIYYPRAGTPEFCVKAAAVKAAMRIQWCPGMRFKMAFETEDSSRISWfMGTISSVQVADPIRWPNSPWRLLQ 378
68899*****************************************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 1.7E-37 | 117 | 229 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 1.11E-28 | 121 | 231 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 4.40E-21 | 124 | 225 | No hit | No description |
| SMART | SM01019 | 6.3E-22 | 125 | 227 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 12.999 | 125 | 227 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 8.7E-22 | 125 | 226 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 1.1E-32 | 295 | 378 | IPR010525 | Auxin response factor |
| PROSITE profile | PS51745 | 19.234 | 620 | 700 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007389 | Biological Process | pattern specification process | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0048829 | Biological Process | root cap development | ||||
| GO:0051301 | Biological Process | cell division | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 706 aa Download sequence Send to blast |
MITLSTEPVK ESMKDADKCL DSQLWHACAG GMVQMPAVNS KVYYFPQGHA EHAQGIVDFG 60 SFPRVPAFIL CRVTAVKFMA DPETDEVFAK IQMVPVRSNE PDSGDDGGMG LGSNGADVQE 120 KPASFAKTLT QSDANNGGGF SVPRYCAETI FPRLDYSADP PVQTVLAKDV HGEVWKFRHI 180 YRGTPRRHLL TTGWSTFVNQ KKLVAGDSIV FLRAENGDLC VGIRRAKRGV GGGHESPSGW 240 NPPTGNCVSA YGGFSVFLRE EENKLMRSNG NGCNLDSGGN MRGRGRVRAD SVIEAATLAA 300 NGQPFEAIYY PRAGTPEFCV KAAAVKAAMR IQWCPGMRFK MAFETEDSSR ISWFMGTISS 360 VQVADPIRWP NSPWRLLQVT WDEPDLLQNV KRVSPWLVEL VSNMPAIHLT PFSPPRKKLC 420 IPQRPDFPIE GQLPTPAFPR NPFGPGSSPL CCFPDSTPAG IQGARHAQFG LSLSDLRLNK 480 LQPGLFHAGL HHLDQATAHS RIPTGLIFGN PAIHDNVSCL LTIGNPSQGT KKLCNGRSPQ 540 LVLFGQPILT EQQISLSNSG DTVSPGVTGN SSLDGNPERT TNVSDGSGSA VHQNGLAENS 600 SCDGFPWYRD RQASELGLET GHCKVFMESE DVGRTLDLSV LGSYEELYGR LADMFGIDKS 660 EMMSHVLYRD AVGAVKQTGD EPFSEFMKTA RRLTILTDSG SDNVGR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldy_A | 8e-73 | 19 | 399 | 18 | 353 | Auxin response factor 1 |
| 4ldy_B | 8e-73 | 19 | 399 | 18 | 353 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00461 | DAP | Transfer from AT4G30080 | Download |
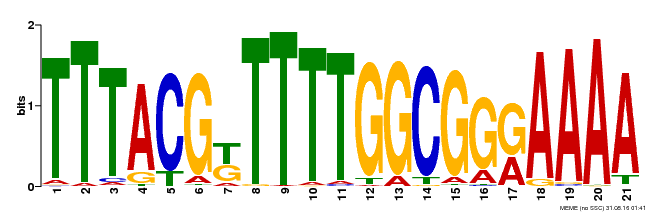
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010934336.2 | 0.0 | auxin response factor 18 | ||||
| Swissprot | Q653H7 | 0.0 | ARFR_ORYSJ; Auxin response factor 18 | ||||
| TrEMBL | A0A2H3XKR6 | 0.0 | A0A2H3XKR6_PHODC; Auxin response factor | ||||
| STRING | XP_008785274.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP516 | 37 | 152 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G30080.1 | 0.0 | auxin response factor 16 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



