 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010934097.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Arecoideae; Cocoseae; Elaeidinae; Elaeis
|
||||||||
| Family | SRS | ||||||||
| Protein Properties | Length: 337aa MW: 35020.7 Da PI: 7.7897 | ||||||||
| Description | SRS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF702 | 224.3 | 2.4e-69 | 98 | 252 | 3 | 153 |
DUF702 3 sgtasCqdCGnqakkdCaheRCRtCCksrgfdCathvkstWvpaakrrerqqqlaaasskaaasa.....aeaaskrkrelkskkqsalsstkls 92
g++sCqdCGnqakkdC h RCRtCCksrgf+C thvkstWvpaakrrerqqqla+as++++ ++ + + skr+re + ++ +t +s
XP_010934097.1 98 GGGKSCQDCGNQAKKDCPHLRCRTCCKSRGFQCPTHVKSTWVPAAKRRERQQQLASASQSHSLRSaavgvSGEPSKRPREICPRL-PTAIATDTS 191
5788**************************************************9998877666568888889999999985555.555667778 PP
DUF702 93 saeskkeletsslPeevsseavfrcvrvssvddgeeelaYqtavsigGhvfkGiLydqGle 153
sa +e+ s+P+evss+avfrcvrvs +d++e+e+aYqta+sigGhvfkGiLydqG++
XP_010934097.1 192 SAGGGGGMEAGSFPAEVSSAAVFRCVRVSPMDEAEDEFAYQTAISIGGHVFKGILYDQGPD 252
888899*****************************************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF05142 | 1.7E-66 | 100 | 252 | IPR007818 | Protein of unknown function DUF702 |
| TIGRFAMs | TIGR01623 | 8.6E-27 | 102 | 144 | IPR006510 | Zinc finger, lateral root primordium type 1 |
| TIGRFAMs | TIGR01624 | 6.5E-25 | 204 | 251 | IPR006511 | Lateral Root Primordium type 1, C-terminal |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009938 | Biological Process | negative regulation of gibberellic acid mediated signaling pathway | ||||
| GO:0010051 | Biological Process | xylem and phloem pattern formation | ||||
| GO:0010252 | Biological Process | auxin homeostasis | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048479 | Biological Process | style development | ||||
| GO:0048480 | Biological Process | stigma development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0046982 | Molecular Function | protein heterodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 337 aa Download sequence Send to blast |
MAGFSLGGSH RQGGEEEEQG GIPPENLFLY SGSGARSEEI PYTRGFELWQ QQQQIQRHQQ 60 HFYPSSALLS FSDEPPGHPP QSGVRGGGGG GGRMSGGGGG KSCQDCGNQA KKDCPHLRCR 120 TCCKSRGFQC PTHVKSTWVP AAKRRERQQQ LASASQSHSL RSAAVGVSGE PSKRPREICP 180 RLPTAIATDT SSAGGGGGME AGSFPAEVSS AAVFRCVRVS PMDEAEDEFA YQTAISIGGH 240 VFKGILYDQG PDGAPFPTTT TPRHIHHGES SSSSSAAAAI TTTAAALSTG LAASTSATAG 300 TTAAASTGFL DPSSLYPAPL SAFMAGTQFF PHHPRPS |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 84 | 92 | RGGGGGGGR |
| 2 | 90 | 98 | GGRMSGGGG |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds DNA on 5'-ACTCTAC-3' and promotes auxin homeostasis-regulating gene expression (e.g. YUC genes), as well as genes affecting stamen development, cell expansion and timing of flowering. Synergistically with other SHI-related proteins, regulates gynoecium, stamen and leaf development in a dose-dependent manner, controlling apical-basal patterning. Promotes style and stigma formation, and influences vascular development during gynoecium development. May also have a role in the formation and/or maintenance of the shoot apical meristem (SAM). Suppressor of the gibberellin (GA) signal transduction. {ECO:0000269|PubMed:10368174, ECO:0000269|PubMed:11706176, ECO:0000269|PubMed:16740146}. | |||||
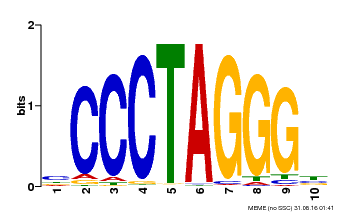
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00613 | PBM | Transfer from AT3G51060 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010934097.1 | 0.0 | protein SHI RELATED SEQUENCE 1-like isoform X1 | ||||
| Swissprot | Q9XGX0 | 6e-70 | SHI_ARATH; Protein SHORT INTERNODES | ||||
| TrEMBL | A0A3Q0HVX0 | 1e-139 | A0A3Q0HVX0_PHODC; protein SHORT INTERNODES-like | ||||
| STRING | XP_008783767.1 | 1e-139 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1354 | 35 | 124 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G51060.1 | 9e-50 | Lateral root primordium (LRP) protein-related | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



