 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010933869.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Arecoideae; Cocoseae; Elaeidinae; Elaeis
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 813aa MW: 87363.8 Da PI: 5.9203 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 67.5 | 1.7e-21 | 125 | 180 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+k+l+L+ rqVk+WFqNrR+++k
XP_010933869.1 125 KKRYHRHTPQQIQELEALFKECPHPDEKQRLELSKRLNLESRQVKFWFQNRRTQMK 180
688999***********************************************999 PP
| |||||||
| 2 | START | 189.7 | 1.4e-59 | 333 | 558 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEE CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla....kaetl 82
ela++a++elvk+a+ +ep+Wv s e ++ +e+ ++f++ + + +ea r++g+v+ ++ lve+l+d + +W +++ + +t+
XP_010933869.1 333 ELALAAMDELVKMAQMDEPLWVPSLegvkETLDDEEYERIFPRCIGmkpvgFVSEATRETGLVIINSSALVETLMDAA-RWADMFPcmiaRSTTT 426
5899*********************9**999**********999889*******************************.**************** PP
EEECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--..-TTSEE-EESSEEEEEEEECTCEEEEEEEE- CS
START 83 evissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe.sssvvRaellpSgiliepksnghskvtwveh 169
evissg g+lqlm+aelq+lsplv+ R++ f+R+++q +g w++vdvS+d +++++ +++++lpSg+++++++ng+skvtwveh
XP_010933869.1 427 EVISSGmggtrnGTLQLMYAELQVLSPLVSvREVNFLRFCKQIAEGAWAVVDVSIDGIRENSSaPTANIKCRRLPSGCVVQDMPNGYSKVTWVEH 521
*************************************************************98766779************************** PP
EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 170 vdlkgrlphwllrslvksglaegaktwvatlqrqcek 206
++++++ +h l+r+l++sgl ga++wvatlqrqce+
XP_010933869.1 522 AEYDEAAVHPLYRPLLRSGLGLGARRWVATLQRQCEC 558
***********************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 5.43E-21 | 114 | 182 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 7.2E-22 | 115 | 184 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 18.074 | 122 | 182 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 3.7E-19 | 123 | 186 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 4.9E-19 | 125 | 180 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.82E-19 | 125 | 183 | No hit | No description |
| PROSITE pattern | PS00027 | 0 | 157 | 180 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 46.117 | 324 | 561 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 2.4E-32 | 327 | 558 | No hit | No description |
| CDD | cd08875 | 1.58E-121 | 330 | 557 | No hit | No description |
| SMART | SM00234 | 3.9E-44 | 333 | 558 | IPR002913 | START domain |
| Pfam | PF01852 | 1.2E-51 | 333 | 558 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 8.43E-23 | 579 | 786 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 813 aa Download sequence Send to blast |
MSFGGLFDGG SGGARVVADF PYGSSAMPAG AISQPRLVSP SLSKPLLSSP GLSLALQTNM 60 EGQGDRSLAS IVGGAGGGGH GGGGGGRDLD AVGTHREDEN ESKSGSENLE GGSGDDIDQE 120 NPRKKKRYHR HTPQQIQELE ALFKECPHPD EKQRLELSKR LNLESRQVKF WFQNRRTQMK 180 TQMERHENTI LKQENDKLRA ENMSIREAMR NPICSNCGGA AVLGEISLEE QHLRIENARL 240 KDELDRVCAL AGKFLGKPIS ALANPLPLPI LNSSLELAVG TTAFAGLGSV PMNLPVVTDF 300 PTGASSPLGS VITPVRPICS MGGVERERTM LLELALAAMD ELVKMAQMDE PLWVPSLEGV 360 KETLDDEEYE RIFPRCIGMK PVGFVSEATR ETGLVIINSS ALVETLMDAA RWADMFPCMI 420 ARSTTTEVIS SGMGGTRNGT LQLMYAELQV LSPLVSVREV NFLRFCKQIA EGAWAVVDVS 480 IDGIRENSSA PTANIKCRRL PSGCVVQDMP NGYSKVTWVE HAEYDEAAVH PLYRPLLRSG 540 LGLGARRWVA TLQRQCECLA ILMSSSLPTR DHTPITASGR RSMLKLAQRM TDNFCAGVCA 600 SSAHKWSKLS LGNIGEDVRV MTRQSVDDPG EPPGVVLSAA TSVWLPVSPM RLFDFLRDER 660 LRSEWDILSN GGPMQEMAHI AKGQDHGNAV SLLRASAVSA NQSSMLILQE TCTDASGSMV 720 VYAPVDIPAM HVVMNGGDSA YVALLPSGFA ILPDGGGGGG GPQKVGGSLL TVAFQILVNS 780 QPTAKLSVES VETVNNLISC TVQKIKAALH CEP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
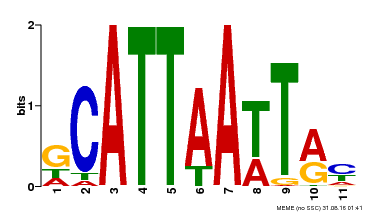
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010933869.1 | 0.0 | homeobox-leucine zipper protein ROC5 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A2H3Y9I9 | 0.0 | A0A2H3Y9I9_PHODC; homeobox-leucine zipper protein ROC5 | ||||
| STRING | XP_008795401.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1203 | 37 | 116 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



