 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010929337.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Arecoideae; Cocoseae; Elaeidinae; Elaeis
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 327aa MW: 34320.1 Da PI: 7.6141 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 56.3 | 4.4e-18 | 243 | 280 | 1 | 36 |
GATA 1 CsnCgttk..TplWRrgpdgnktLCnaCGlyyrkkglk 36
C++Cgt++ Tp++Rrgpdg++tLCnaCGl+++ kg++
XP_010929337.1 243 CHHCGTSSkaTPMMRRGPDGPRTLCNACGLMWANKGTM 280
***********************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00979 | 2.7E-12 | 107 | 142 | IPR010399 | Tify domain |
| PROSITE profile | PS51320 | 14.892 | 107 | 142 | IPR010399 | Tify domain |
| Pfam | PF06200 | 6.2E-13 | 110 | 141 | IPR010399 | Tify domain |
| Pfam | PF06203 | 6.3E-16 | 168 | 210 | IPR010402 | CCT domain |
| PROSITE profile | PS51017 | 13.304 | 168 | 210 | IPR010402 | CCT domain |
| SMART | SM00401 | 7.6E-13 | 237 | 290 | IPR000679 | Zinc finger, GATA-type |
| PROSITE profile | PS50114 | 10 | 237 | 299 | IPR000679 | Zinc finger, GATA-type |
| SuperFamily | SSF57716 | 4.75E-13 | 241 | 294 | No hit | No description |
| Gene3D | G3DSA:3.30.50.10 | 2.9E-16 | 242 | 284 | IPR013088 | Zinc finger, NHR/GATA-type |
| CDD | cd00202 | 1.31E-16 | 242 | 285 | No hit | No description |
| Pfam | PF00320 | 1.0E-15 | 243 | 280 | IPR000679 | Zinc finger, GATA-type |
| PROSITE pattern | PS00344 | 0 | 243 | 270 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 327 aa Download sequence Send to blast |
MSQSPTKPAD VKPLAGGRYT ARHPAPAHPA VPLPGRIDDA SGSGGGGGEG QLIVAADAQA 60 IQQYAADGGA GDGMEEDHDG VAEGEGMEGD VPSDHANLGD PHGVVAPQVG ANQLTLSFQG 120 EVYVFDSVSP EKVQAVLLLL GGREINTGLA SGPSSSNPYN KRSNFPQRVA SLMRFREKRK 180 ERNFDKKIRY NVRKEVALRM QRNRGQFTSS KSKPEDAAAG ITSLDVSQRW GSVEGRPESA 240 ATCHHCGTSS KATPMMRRGP DGPRTLCNAC GLMWANKGTM RDLSKNSPAA TQNALLETKE 300 GDGNGAAVGE QQQQPLPITA NGHGSSS |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that specifically binds 5'-GATA-3' or 5'-GAT-3' motifs within gene promoters. {ECO:0000250|UniProtKB:Q8LAU9}. | |||||
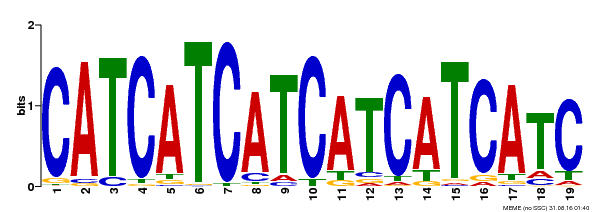
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00198 | DAP | Transfer from AT1G51600 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By abscisic acid (ABA), and drought and salt stresses. Down-regulated by jasmonate and wounding. {ECO:0000269|PubMed:19618278}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010929337.1 | 0.0 | GATA transcription factor 20 | ||||
| Swissprot | Q5Z4U5 | 1e-112 | GAT20_ORYSJ; GATA transcription factor 20 | ||||
| TrEMBL | A0A2H3ZBR9 | 0.0 | A0A2H3ZBR9_PHODC; GATA transcription factor 20-like | ||||
| STRING | XP_008811434.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4325 | 38 | 68 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G21175.2 | 3e-83 | ZIM-like 1 | ||||



