 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010928389.1 | ||||||||
| Common Name | SREBP | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Arecoideae; Cocoseae; Elaeidinae; Elaeis
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 372aa MW: 40387.7 Da PI: 8.9999 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 63 | 5.8e-20 | 16 | 61 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+g+W++eEde l ++v+++G+++W++I++ ++ gR++k+c++rw +
XP_010928389.1 16 KGPWSPEEDEALQKLVQKHGPRNWSLISKSIP-GRSGKSCRLRWCNQ 61
79******************************.***********985 PP
| |||||||
| 2 | Myb_DNA-binding | 54.8 | 2.1e-17 | 70 | 112 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
++T+eEde +++a++++G++ W+tIar + gRt++ +k++w++
XP_010928389.1 70 PFTAEEDETIIRAHRRFGNK-WATIARLLS-GRTDNAIKNHWNST 112
89******************.*********.***********986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 28.066 | 11 | 66 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.25E-32 | 13 | 109 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 6.4E-18 | 15 | 64 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.8E-19 | 16 | 61 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 6.2E-27 | 17 | 69 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 8.15E-17 | 18 | 60 | No hit | No description |
| SMART | SM00717 | 7.6E-15 | 67 | 115 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS51294 | 20.613 | 68 | 117 | IPR017930 | Myb domain |
| CDD | cd00167 | 5.08E-12 | 70 | 113 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 3.0E-23 | 70 | 116 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 2.1E-14 | 70 | 112 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 372 aa Download sequence Send to blast |
MAASISNRGK DMDRIKGPWS PEEDEALQKL VQKHGPRNWS LISKSIPGRS GKSCRLRWCN 60 QLSPQVEHRP FTAEEDETII RAHRRFGNKW ATIARLLSGR TDNAIKNHWN STLKRKYSSA 120 TVAPAVDDVG LPHQQYDAAL VAADDAAVRP LKRSSSVGPI LSSAGGLCLS PGSPSGSDLS 180 DSSRHSHPMF SPAAAAAVLP SASHIYRPVP RTGGIVRPAS SPHHHEATVV AAPAINTKND 240 DPVTSLSLSL PGSDKPEASD LHRTNQNQLH LLAAPLSPGQ AMPLHPPTPV AHPPPPMACP 300 STADEQRREP DEERRPAFPF SAEFLAVMQE MIRNEVRSYM SGLEQSGMMC MQPPQESIHN 360 AVIKRIGITK IE |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1mse_C | 2e-41 | 15 | 116 | 3 | 104 | C-Myb DNA-Binding Domain |
| 1msf_C | 2e-41 | 15 | 116 | 3 | 104 | C-Myb DNA-Binding Domain |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00476 | DAP | Transfer from AT4G37260 | Download |
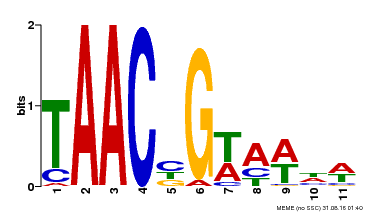
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010928389.1 | 0.0 | transcription factor MYB44 | ||||
| TrEMBL | K0A224 | 0.0 | K0A224_ELAGV; Sucrose responsive element-binding protein | ||||
| STRING | XP_008799576.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP789 | 38 | 154 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G67300.1 | 2e-69 | myb domain protein r1 | ||||



