 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010923453.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Arecoideae; Cocoseae; Elaeidinae; Elaeis
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 367aa MW: 40112.3 Da PI: 10.0378 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 39.7 | 1.2e-12 | 52 | 100 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s++kGV + +grW A+I++ ++r++lg+f + eAa+a++ a+++++g
XP_010923453.1 52 SHFKGVVPQP-NGRWGAQIYEK-----HQRVWLGTFNEESEAARAYDVAAQRFRG 100
789***9788.8*********3.....5**********88*************98 PP
| |||||||
| 2 | B3 | 102.8 | 1.9e-32 | 191 | 291 | 1 | 96 |
EEEE-..-HHHHTT-EE--HHH.HTT.....---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.....ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsef 90
f+k+ltpsdv+k++rlv+pk++ae+h g +++ l +ed+ g++W+++++y+++s++yvltkGW++Fvk++ Lk+gD+v+F+++ +++
XP_010923453.1 191 FDKALTPSDVGKLNRLVIPKQHAEKHfpiqiAGAAGKGVLLNFEDSCGKVWRFRYSYWNSSQSYVLTKGWSRFVKEKSLKAGDVVTFQRSTGQDK 285
89****************************97777899**************************************************8766777 PP
..EEEE CS
B3 91 elvvkv 96
+l++ +
XP_010923453.1 286 QLFIDW 291
777766 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 9.96E-24 | 52 | 107 | No hit | No description |
| SuperFamily | SSF54171 | 1.05E-16 | 52 | 108 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 4.8E-8 | 52 | 100 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 4.0E-20 | 53 | 108 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 4.9E-28 | 53 | 114 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.612 | 53 | 108 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 3.4E-42 | 186 | 295 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 1.7E-31 | 188 | 292 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.84E-28 | 189 | 280 | No hit | No description |
| Pfam | PF02362 | 1.3E-29 | 191 | 292 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 2.1E-23 | 191 | 295 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 14.113 | 191 | 294 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 367 aa Download sequence Send to blast |
MDSRCIEDVC SIRTEKRPPP GPESLQRVGS GASVVLDPEV GVEAESRKLP SSHFKGVVPQ 60 PNGRWGAQIY EKHQRVWLGT FNEESEAARA YDVAAQRFRG RDAVTNFKPF SETDDDDAAE 120 LSFLASHSKA EIVDMLRKHT YHDEVRQSKR AFGAGKGSDR RSMPAGGYGG GLGLRFLAGS 180 GGGGAQREHL FDKALTPSDV GKLNRLVIPK QHAEKHFPIQ IAGAAGKGVL LNFEDSCGKV 240 WRFRYSYWNS SQSYVLTKGW SRFVKEKSLK AGDVVTFQRS TGQDKQLFID WKPRAAGGCA 300 EARPVQGIRL FGVNLFKIPA VAVGGAGLDG NGGVDHVVGC SRKGIRGMDH QLMPSPQQFK 360 KQKTEAL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wid_A | 1e-56 | 188 | 307 | 11 | 130 | DNA-binding protein RAV1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional repressor of flowering time on long day plants. Acts directly on FT expression by binding 5'-CAACA-3' and 5'-CACCTG-3 sequences. Functionally redundant with TEM2. {ECO:0000269|PubMed:18718758}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00024 | PBM | Transfer from AT1G25560 | Download |
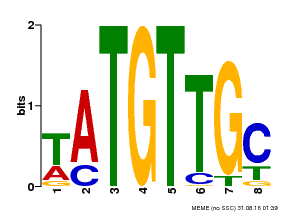
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Expressed with a circadian rhythm showing a peak at dawn. {ECO:0000269|PubMed:18718758}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010923453.1 | 0.0 | AP2/ERF and B3 domain-containing transcription factor RAV1 | ||||
| Swissprot | Q9C6M5 | 1e-131 | RAVL1_ARATH; AP2/ERF and B3 domain-containing transcription repressor TEM1 | ||||
| TrEMBL | A0A2H3Y084 | 0.0 | A0A2H3Y084_PHODC; AP2/ERF and B3 domain-containing transcription factor RAV1 | ||||
| STRING | XP_008791228.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP456 | 38 | 203 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G13260.1 | 1e-130 | related to ABI3/VP1 1 | ||||



