 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010922397.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Arecoideae; Cocoseae; Elaeidinae; Elaeis
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 187aa MW: 21560.9 Da PI: 4.7258 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 52 | 1.6e-16 | 2 | 43 | 5 | 48 |
-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 5 TteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
T++E+ l+++++ ++G++ W++Iar+++ gRt++++k++w++++
XP_010922397.1 2 TPQEERLILELHSRWGNR-WSRIARKLP-GRTDNEIKNYWRTHM 43
9*****************.*********.************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00167 | 1.14E-11 | 1 | 43 | No hit | No description |
| PROSITE profile | PS51294 | 24.852 | 1 | 47 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.6E-11 | 1 | 45 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 9.36E-13 | 2 | 49 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.2E-20 | 2 | 44 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 9.4E-15 | 2 | 42 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 187 aa Download sequence Send to blast |
MTPQEERLIL ELHSRWGNRW SRIARKLPGR TDNEIKNYWR THMRKKAQER KKNALSSSSS 60 SSSLTDNSAS VSEPPSDAGL EQHELNGGSG CCCMAAGFVE NEEGVKGYSM DQIWNEIIAT 120 PEAVSELSFG EWKNDFFDTS WPPMPSPMWE YSSESPWKMD YEEFKMLPSL DDLLLFSNYQ 180 HGREFYN |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. {ECO:0000305}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00600 | PBM | Transfer from AT5G59780 | Download |
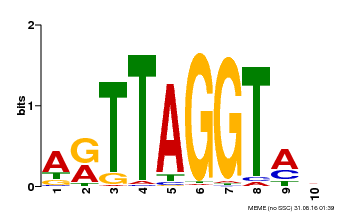
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010922397.1 | 1e-138 | myb-related protein MYBAS2 isoform X2 | ||||
| Swissprot | Q4JL76 | 7e-42 | MYBA2_ORYSJ; Myb-related protein MYBAS2 | ||||
| TrEMBL | A0A2H3XSU6 | 3e-87 | A0A2H3XSU6_PHODC; myb-related protein MYBAS2 isoform X1 | ||||
| TrEMBL | A0A3Q0I2V1 | 3e-88 | A0A3Q0I2V1_PHODC; myb-related protein MYBAS2 isoform X2 | ||||
| STRING | XP_008788078.1 | 5e-88 | (Phoenix dactylifera) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G59780.1 | 1e-37 | myb domain protein 59 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



